Non-invasive biomarkers in pancreatic cancer diagnosis: what we need versus what we have
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the predominant histologic type of pancreatic cancer (PC) and has one of the worse prognoses of all types of cancer leading to 227,000 deaths annually worldwide. It is currently the fourth leading cause of cancer-related death in the world. Incidence of PC in the US and most of developed countries continue to rise with forecast claiming that it will be the second most fatal cancer in the US by 2020 (1).
The majority of patients with PC progress to either locally advanced or metastatic disease in the asymptomatic phase, as many as 80% presents late with metastasis at diagnosis (2). Metastasis is the most common cause of death in PC patients. Without possibility for resection, survival at stage 4 is <5%. When diagnosed at stage 1, 5-year survival is around 20%. However, if detection would be possible before patient present clinical symptoms, survivability could reach 75% after 5 years.
Although for many years the question of whether the dismal prognosis of PC compared with other types of tumors would be due to late diagnosis or early dissemination of malignant cells, several studies provided novel insights. Yachida et al. (3) performed experiments indicating that at least 5 years are required for the acquisition of metastatic ability after a parental tumoral but non-metastatic founder cell is detected. These data define a very important broad time window in which there are opportunities for early detection that could prevent deaths from PC metastatic disease. This is generally accepted nowadays that diagnose PC at early stages is needed since it will improve survival.
PDAC are generally developed from three precursor lesions: pancreatic intraepithelial neoplasia (PanIN) lesions, intraductal papillary mucinous neoplasms (IPMN) and mucinous cystic neoplasms (MCN) (4). While PanIN lesions are microscopic and not detectable by image techniques, cystic premalignant lesions (IPMN and MCN) are seen by modalities such as endoscopic ultrasound (EUS), magnetic resonance with cholangiopancreatography and computerized tomography scan (CT scan). However, the large majority of these cystic lesions do not progress to cancer, their incidental detection often results in difficulties regarding clinical management in terms of surgery recommendation.
Diagnosis of PC is nowadays very challenging since patients present with non-specific symptoms leading to delay in correct diagnosis. Additionally, and very importantly, results obtained from imaging techniques are not conclusive and often of ambiguous relevance. In fact, while sometimes pancreatic mass are indistinguishable from chronic pancreatitis or benign pancreatic cysts when biopsy is obtained from the lesion, pathological results can be inconclusive. Cytological examination of sample obtained by endoscopic ultrasound-guided fine needle aspiration (EUS-FNA) may be non-precise due to sampling difficulties, inflammation coexistence… etc. (5). For these reasons, many clinical and basic investigators are trying to identify biomarkers that could support gastroenterologists and pathologists in PC diagnosis.
Biomarkers needed
General population screening is not advisable due that the overall lifetime risk of developing PC is relatively low, close to 1%, and does not meet some of the criteria established by the World Health Organization (http://www.who.int/cancer/detection/variouscancer/en). Additionally, since there is limited evidence about accuracy of PC screening tests, acceptability, cost and availability on whom to treat on the basis of screening results, this approach is not currently advisable for average risk subjects.
By contrast, only individuals with a significant increased risk of develop PC could opt to screening test. The International Cancer of the Pancreas Screening Consortium (CAPS) (5) have recently stated that screening is indicated for: (I) individuals with “familiar pancreatic cancer” (FPC), with two or more blood relatives affected by PDAC, of whom at least one first degree relative (FDR); and (II) genetic syndromes such as Peutz-Jeghers syndrome and p16 [familial atypical multiple mole melanoma syndrome (FAMMM syndrome)], BRCA2, PALB and hereditary non-polyposis colorectal cancer mutation carriers if one FDR or two family members are affected by PC. However, for this high risk considered patients, there is not consensus about screening modality used but most commonly magnetic resonance imaging (MRI) or EUS have been proposed. There is not evidence of blood biomarkers being advised for this purpose. In fact, the diagnostic utility of CA19-9 in high risk individuals has been fully investigated but available data indicate that this biomarker does not seem helpful in this population (6).
Considering the dismal prognosis of PC due to late diagnosis and accepting that current techniques make screening for general population not advisable and sometimes uncertain for high-risk patients, there is an urgent need to find new biomarkers that could cover several unmet clinical needs: (I) screening of high risk individuals; and (II) confirmation in differential diagnosis for medium risk individuals.
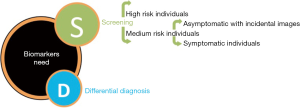
For medium risk individuals, a clinically useful biomarker should (I) allow early diagnosis of patients with incidental image findings, at its earliest forms (ideally asymptomatic, before symptoms starts) (II) helping in a definitive diagnosis by distinguish PC from diseases with similar non specific symptoms such as pancreatitis or benign pancreatic cyst (differential diagnosis) or those with strong but not definitive image suspicion (confirmatory diagnosis) (Figure 1).
Considerations for biomarkers development
A useful biomarker with applicability for early diagnosis should be minimally or non-invasive and have high sensitivity, high specificity and capacity to discriminate low-grade dysplasia from high-grade dysplasia and cancer.
Minimally-invasive or non-invasive biomarkers are required to encourage use for clinicians and compliance in patients. Only non-invasive tests are practical enough and will be able to maximize access. To obtain these properties, biomarkers in blood, urine, saliva, feces or pancreatic juice samples must be investigated.
Sensitive biomarkers are required to identify correctly those patients who have PC. However, high sensitive biomarkers with low specificity results in a larger number of false positive results leading to unnecessary, expensive and not affordable procedures. Only cost-effective and inexpensive biomarkers will be widely distributable being finally accessible to health-care systems and patients.
By the development of non invasive biomarkers for early diagnosis an impact will be obtained for: (I) those patients in whom clinically suspected PC is part of differential diagnosis; (II) those patients with strong clinical and image suspicion of PC; (III) patients with medium or high risk of PC development.
To achieve these aims, performance characteristics including non-invasiveness, accuracy (sensitive and specific) and cost-effectiveness biomarker must be identified (Table 1).

Full table
Conventional approaches
Carbohydrate antigen 19-9 (CA19-9) and CA19-9 combinatory panels
CA19-9 is the only biomarker widely used for the management of PC so far (7). However, due to its important limitations CA19-19 is an unreliable screening biomarker being restricted to detection of tumor recurrence after surgical resection (8-10). CA19-9 shows lack of expression in ~5% of the population (Caucasians lacking the Lewis blood group antigen) while an elevation can be observed in related diseases including chronic pancreatitis and obstructive jaundice (9,11). CA19-9 has a variable sensitivity of ~85% and specificity of ~85% for the detection of PC (12) but the low prevalence of the disease makes this biomarker not applicable for screening and not relevant to confirmatory/differential diagnosis.
Although not specific enough, CA19-9 seem to be most widely used biomarker in PC and consequently, multiple investigations have combined CA19-9 with other markers (Including factors produced by the tumor or molecules from systemic response to the growing tumor or inflammatory reactants) in order to improve accuracy (13). For instance the combination of CA19-9, IGF-1 and albumin resulted in a sensitivity of 93.6% and specificity of 95% (13) when differentiating pancreatitis from PC. Serum cytokines as biomarker panel including CA19-9 showed a sensitivity 85.7% and specificity of 92.3% to distinguish PC from healthy controls and sensitivity of 98%, specificity of 96.4% to distinguish PC from chronic pancreatitis (14). Also, Brand et al. (15) reported that the combination of CA19-9, ICAM-1, and OPG discriminated PC patients from healthy controls with a sensitivity/specificity of 88/90%, while the panel of CA 19-9, carcinoembryonic antigen (CEA), and TIMP-1 discriminated PC patients from benign subjects with a sensitivity/specificity of 76/90%. A panel consisting of carbohydrate antigen CA19-9, albumin, C-reactive protein and interleukin 28 demonstrated sensitivity was 99.39% for all-stages, 96.10% for early-stage and 98.80% for advanced-stage PDAC at 90% specificity when discriminating between PDAC and healthy individuals.
However, all these studies analyzing a panel of biomarker should be validated in larger cohorts. Independent validation sets are required to further corroborate obtained data since it is very common that training set values are higher than those obtained in validations sets. Due to these reasons, the lack of reproducibility leads to non-clinical applicability.
Carcinoembryonic antigen (CEA)
CEA was found in gastrointestinal tissue during fetal development and colorectal carcinoma, and was later purpose to diagnose PC. Several meta-analysis showed that CA19-9 have better performance than CEA. Nevertheless, combination of these two biomarkers have been very popular in panels (16). Nowadays, this biomarker is not used to diagnose PC as the protein is not produced by all pancreatic tumors and has lower sensitivity than CA19-9 (17).
Novel approaches
miRNAs
miRNAs are non-coding RNAs that regulates gene expression and have been recognized as deregulated in oncogenesis (18,19). miRNAs are very stable in blood because they are usually bound to Argonaute, a protein that protects them from RNase degradation (20,21). Several authors reported that miRNAs are deregulated in pancreatic diseases being able to discriminate PC from pancreatitis, pancreatic precursor lesions and healthy individuals (22,23) and precursor lesions such as IPMN with malignant potential (24) being present in several types of samples.
Schultz et al. (23) from the Danish BIOPAC study identified two miRNA panels consisting in groups of four biomarkers that distinguish PC from healthy controls (miR-145, miR-150, miR-223, miR-636) and ten mi-RNAs (miR-26b, miR-34a, miR-122, miR-126*, miR-145, miR-150, miR-223, miR-505, miR-636, miR-885.5p). Li et al. (19) showed miR-1290 as biomarker able to distinguish early PC from healthy subjects. Several authors found that miR-216 and miR-217 are down-regulated in PC while miR-143, miR145, miR-146, miR-148, miR-150, miR155, miR-196a, mir-196b, miR-210, miR-222, miR-223 and miR-31 are up-regulated in PC (25-28). Komatsu et al. suggested that plasma miR-223 might be a clinically useful biomarker for diagnosing PC, and predicting malignant potential of IPMN and the invasiveness of PC (24). Hernandez et al. reviewed the current knowledge of miRNA in PC and its precursor lesions concluding that miR-21, miR-155, miR-196 and miR-210 are dysregulated in tissue, serum, cyst fluid and stool of PDAC patients. miR-21, miR-155 and miR-196 are dysregulated in PanIN and IPMN lesions (29).
However, one of the main challenges in miRNAs evaluation is the difficulty to detect accurately the amount of these biomarkers due they are short molecules that could be bound to proteins or even integrated in vesicles. Currently, qRT-PCR, in situ hybridization, microarray and next-generation sequencing assays are the most frequent techniques used for miRNA evaluation (5). Still, time is needed to make possible some of these techniques are part of the clinical routine allowing part of diagnostic laboratories.
In definitive, miRNAs have been profiled as one of the most popular non-invasive biomarker in early detection of PC and numerous of these molecules are shown to be dysregulated while and after PC development. Validations in larger cohorts and consensus regarding definitive panels are need before clinical applications would be real for patients.
Methylation biomarkers
Several studies have focused on the methylation status as biomarker for PDAC early detection or differential diagnosis. Methylation patterns have been observed in pancreatic juice (30,31) and stool (32) from patients with PC and precursor lesions.
Recently, investigators from Mayo Clinic speculated that novel methylation markers that distinguish PC from benign controls are detected in pancreatic juice. Kisiel et al. (33) described a panel of methylated biomarkers CD1D, KCNK12, CLEC11A, NDRG4, IKZF1, PKRCB and KRAS resulting in 75% sensitivity and 95% specificity comparing PC to normal pancreas and pancreatitis. Presently, a larger clinical study is being performed to assess this accuracy. Similarly to Septin-9 methylation experience as screening biomarker in colon cancer, investigators expect rapid validation and translation to the clinic.
Other methylation biomarkers associated to disease are ppENK, cyclin D2, sparc-7, Osteonectin and TSLC1 but not further validation and clinical application has been found yet (34-37).
Circulating tumor DNA (ctDNA) and circulating tumor cells (CTC)
Liquid biopsy is one of the most recent approaches in oncology mostly for relapse detection in patients receiving treatment and it has been also explored in PC (38). In this, ctDNA and CTC are the most promising blood biomarkers, often starting to be used with prognostic or relapse purposes.
ctDNA measurements on KRAS mutations seems to be a marker for monitoring treatment efficacy and PC disease progression instead of having a diagnostic value (39).
CTC are tumor cell that have the ability to enter the circulatory system and are ultimately responsible for metastasis development. However, CTC are extremely rare in PC and both enrichment and isolation has been very difficult so far.
Recent data by Rhim et al. (40,41) suggested that circulating tumor cells could be detectable before primary tumor can be visualized by current image technique. Probed in mouse models but also in humans (where CTC were detected in 73% of PDAC patients and undetectable in healthy subjects), confirmation in larger studies is now required. Difficulties for CTC identification and isolation are one of the principal issues in the scientific community when working with this novel and completely well defined entity. To demonstrate applicability technical and methodological issues required to be clarified.
Some critics say that neither CTC nor ctDNA can be used for PC screening or diagnosis, because of limited sensitivity. However, CTC as seed of metastasis are prognostic biomarkers in non-metastatic diseases and ctDNA might be accurate for relapse prediction and therapy management (42).
Molecules in exosomes
Exosomes are secreted membrane enclosed vesicles formed during the inward budding of endosomes and contains nucleic acid and proteins (43,44). They are secreted by cells, including tumoral cells and circulate in the blood exosomes playing key roles in cell-cell communications, regulating diverse biological processes. Kahlert et al. previously demonstrated that the DNA in the circulation is mainly associated with exosomes (45).
Melo et al. presented a study where exosomes from PC patients express higher levels of GPC1 than healthy subjects (46). Exosomes positive for GPC1 may serve as a potential non-invasive diagnostic and screening tool to detect early stages of PC to facilitate possible curative surgical therapy. Ding et al. reveled that PC-derived exosomes transfer miRNAs to dendritic cells via miR-212 to induce immunotolerance (47).
The isolation of cancer exosomes from patients remains a challenge due to the lack of specific markers that can distinguish cancer from non-cancer exosomes and time-consuming technology required not yet implantable in clinical routine.
Conclusions
Discouraging statistics about 5-year survival after diagnosis have remained fairly consistent for the last decades with no improvement in the prognosis of PC patients. PC is probably the most lethal tumor and by 2020 it will be the second cause of cancer death. It is well-established now that only earliest forms of PC are a curable disease but it has to be detected before symptoms starts. Although several strategies have been proposed, the achievement of detection at curable phase demands screening intervention for early detection and differential diagnosis. Similarly to what occurs in colorectal cancer, development through adenoma-cancer sequence, an increasing number of scientific evidences show that progression from premalignant lesions to cancer also exists in PC.
Unfortunately no successful strategy has been concluded as effective approach and now, non-invasive biomarkers are the hope. Multiple translational research studies have explored minimally or non-invasive biomarkers in biofluids-blood, urine, stool, saliva or pancreatic juice, although diagnostic performance has not been further validated. These include overexpressed and under-expressed microRNAs (25), mutations and other genetic alterations, epigenetic changes such as methylation (48), modification of secreted proteins such as mucins (49) or most recently the detection of free nucleid acids such as ctDNA (50), circulating pancreatic cells (CPCs) (51) or cancer stem cells (CSCs) (52). However, the development achieved so far in science and technology has not resulted in improved survival for patients. Consequently, experts around the world try to fill the gaps in collaborative studies, meetings, conferences and work groups in order to find reliable test for early detection of PC (53).
The failure of single non-invasive biomarker can be explained by the complex biology and heterogeneity of PC and some investigators recommend examining the use of a combination of different biomarkers demonstrating heterogeneous pathophysiology (13).
Nowadays no biomarker, alone or in combination, has been superior to CA19-9 in sensitivity and specificity. Still when lesions are not differentiable by image, molecular approaches will constitute the only solution for diagnostic improvement.
Although the number of novel biomarkers for early diagnosis of PC has been increasing during the last couple of years, no molecular signature has been fully validated to be implemented in clinical routine. Under the uncertain future, miRNAs profiling and methylation status seem to be the most promising biomarkers. However, good results in larger validations are urgently needed before application. Industry efforts through biotech and pharmaceutical companies are urgently required to further validate what basis science established as promising. Clinical studies and trials are important to demonstrate accuracy after preliminary data from academy.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Siegel R, Ma J, Zou Z, et al. Cancer statistics, 2014. CA Cancer J Clin 2014;64:9-29. [Crossref] [PubMed]
- Hidalgo M. Pancreatic cancer. N Engl J Med 2010;362:1605-17. [Crossref] [PubMed]
- Yachida S, Jones S, Bozic I, et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 2010;467:1114-7. [Crossref] [PubMed]
- Klöppel G, Basturk O, Schlitter AM, et al. Intraductal neoplasms of the pancreas. Semin Diagn Pathol 2014;31:452-66. [Crossref] [PubMed]
- Canto MI, Harinck F, Hruban RH, et al. International Cancer of the Pancreas Screening (CAPS) Consortium summit on the management of patients with increased risk for familial pancreatic cancer. Gut 2013;62:339-47. [Crossref] [PubMed]
- Capurso G, Signoretti M, Valente R, et al. Methods and outcomes of screening for pancreatic adenocarcinoma in high-risk individuals. World J Gastrointest Endosc 2015;7:833-42. [Crossref] [PubMed]
- Wong D, Ko AH, Hwang J, et al. Serum CA19-9 decline compared to radiographic response as a surrogate for clinical outcomes in patients with metastatic pancreatic cancer receiving chemotherapy. Pancreas 2008;37:269-74. [Crossref] [PubMed]
- Ballehaninna UK, Chamberlain RS. The clinical utility of serum CA 19-9 in the diagnosis, prognosis and management of pancreatic adenocarcinoma: An evidence based appraisal. J Gastrointest Oncol 2012;3:105-19. [PubMed]
- Locker GY, Hamilton S, Harris J, et al. ASCO 2006 update of recommendations for the use of tumor markers in gastrointestinal cancer. J Clin Oncol 2006;24:5313-27. [Crossref] [PubMed]
- Berger AC, Garcia M Jr, Hoffman JP, et al. Postresection CA 19-9 predicts overall survival in patients with pancreatic cancer treated with adjuvant chemoradiation: a prospective validation by RTOG 9704. J Clin Oncol 2008;26:5918-22. [Crossref] [PubMed]
- Marrelli D, Caruso S, Pedrazzani C, et al. CA19-9 serum levels in obstructive jaundice: clinical value in benign and malignant conditions. Am J Surg 2009;198:333-9. [Crossref] [PubMed]
- Huang Z, Liu F. Diagnostic value of serum carbohydrate antigen 19-9 in pancreatic cancer: a meta-analysis. Tumour Biol 2014;35:7459-65. [Crossref] [PubMed]
- Ferri MJ, Saez M, Figueras J, et al. Improved Pancreatic Adenocarcinoma Diagnosis in Jaundiced and Non-Jaundiced Pancreatic Adenocarcinoma Patients through the Combination of Routine Clinical Markers Associated to Pancreatic Adenocarcinoma Pathophysiology. PLoS One 2016;11:e0147214. [Crossref] [PubMed]
- Zeh HJ, Winikoff S, Landsittel DP, et al. Multianalyte profiling of serum cytokines for detection of pancreatic cancer. Cancer Biomark 2005;1:259-69. [PubMed]
- Brand RE, Nolen BM, Zeh HJ, et al. Serum biomarker panels for the detection of pancreatic cancer. Clin Cancer Res 2011;17:805-16. [Crossref] [PubMed]
- Zhang Y, Yang J, Li H, et al. Tumor markers CA19-9, CA242 and CEA in the diagnosis of pancreatic cancer: a meta-analysis. Int J Clin Exp Med 2015;8:11683-91. [PubMed]
- Duraker N, Hot S, Polat Y, et al. CEA, CA 19-9, and CA 125 in the differential diagnosis of benign and malignant pancreatic diseases with or without jaundice. J Surg Oncol 2007;95:142-7. [Crossref] [PubMed]
- Iorio MV, Croce CM. MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol 2009;27:5848-56. [Crossref] [PubMed]
- Li A, Yu J, Kim H, et al. MicroRNA array analysis finds elevated serum miR-1290 accurately distinguishes patients with low-stage pancreatic cancer from healthy and disease controls. Clin Cancer Res 2013;19:3600-10. [Crossref] [PubMed]
- Fujita N, Noda Y, Kobayashi G, et al. Endoscopic approach to early diagnosis of pancreatic cancer. Pancreas 2004;28:279-81. [Crossref] [PubMed]
- Wakatsuki T, Irisawa A, Bhutani MS, et al. Comparative study of diagnostic value of cytologic sampling by endoscopic ultrasonography-guided fine-needle aspiration and that by endoscopic retrograde pancreatography for the management of pancreatic mass without biliary stricture. J Gastroenterol Hepatol 2005;20:1707-11. [Crossref] [PubMed]
- Bauer AS, Keller A, Costello E, et al. Diagnosis of pancreatic ductal adenocarcinoma and chronic pancreatitis by measurement of microRNA abundance in blood and tissue. PLoS One 2012;7:e34151. [Crossref] [PubMed]
- Schultz NA, Dehlendorff C, Jensen BV, et al. MicroRNA biomarkers in whole blood for detection of pancreatic cancer. JAMA 2014;311:392-404. [Crossref] [PubMed]
- Komatsu S, Ichikawa D, Miyamae M, et al. Malignant potential in pancreatic neoplasm; new insights provided by circulating miR-223 in plasma. Expert Opin Biol Ther 2015;15:773-85. [Crossref] [PubMed]
- Bloomston M, Frankel WL, Petrocca F, et al. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 2007;297:1901-8. [Crossref] [PubMed]
- Szafranska AE, Davison TS, John J, et al. MicroRNA expression alterations are linked to tumorigenesis and non-neoplastic processes in pancreatic ductal adenocarcinoma. Oncogene 2007;26:4442-52. [Crossref] [PubMed]
- Hanoun N, Delpu Y, Suriawinata AA, et al. The silencing of microRNA 148a production by DNA hypermethylation is an early event in pancreatic carcinogenesis. Clin Chem 2010;56:1107-18. [Crossref] [PubMed]
- Liffers ST, Munding JB, Vogt M, et al. MicroRNA-148a is down-regulated in human pancreatic ductal adenocarcinomas and regulates cell survival by targeting CDC25B. Lab Invest 2011;91:1472-9. [Crossref] [PubMed]
- Hernandez YG, Lucas AL. MicroRNA in pancreatic ductal adenocarcinoma and its precursor lesions. World J Gastrointest Oncol 2016;8:18-29. [Crossref] [PubMed]
- Matsubayashi H, Canto M, Sato N, et al. DNA methylation alterations in the pancreatic juice of patients with suspected pancreatic disease. Cancer Res 2006;66:1208-17. [Crossref] [PubMed]
- Kato N, Yamamoto H, Adachi Y, et al. Cancer detection by ubiquitin carboxyl-terminal esterase L1 methylation in pancreatobiliary fluids. World J Gastroenterol 2013;19:1718-27. [Crossref] [PubMed]
- Kisiel JB, Yab TC, Taylor WR, et al. Stool DNA testing for the detection of pancreatic cancer: assessment of methylation marker candidates. Cancer 2012;118:2623-31. [Crossref] [PubMed]
- Kisiel JB, Raimondo M, Taylor WR, et al. New DNA Methylation Markers for Pancreatic Cancer: Discovery, Tissue Validation, and Pilot Testing in Pancreatic Juice. Clin Cancer Res 2015;21:4473-81. [Crossref] [PubMed]
- Fukushima N, Walter KM, Uek T, et al. Diagnosing pancreatic cancer using methylation specific PCR analysis of pancreatic juice. Cancer Biol Ther 2003;2:78-83. [Crossref] [PubMed]
- Ueki T, Walter KM, Skinner H, et al. Aberrant CpG island methylation in cancer cell lines arises in the primary cancers from which they were derived. Oncogene 2002;21:2114-7. [Crossref] [PubMed]
- Matsubayashi H, Sato N, Fukushima N, et al. Methylation of cyclin D2 is observed frequently in pancreatic cancer but is also an age-related phenomenon in gastrointestinal tissues. Clin Cancer Res 2003;9:1446-52. [PubMed]
- Sato N, Fukushima N, Maehara N, et al. SPARC/osteonectin is a frequent target for aberrant methylation in pancreatic adenocarcinoma and a mediator of tumor-stromal interactions. Oncogene 2003;22:5021-30. [Crossref] [PubMed]
- Kinugasa H, Nouso K, Miyahara K, et al. Detection of K-ras gene mutation by liquid biopsy in patients with pancreatic cancer. Cancer 2015;121:2271-80. [Crossref] [PubMed]
- Tjensvoll K, Lapin M, Buhl T, et al. Clinical relevance of circulating KRAS mutated DNA in plasma from patients with advanced pancreatic cancer. Mol Oncol 2015. [Epub ahead of print]. [PubMed]
- Rhim AD, Thege FI, Santana SM, et al. Detection of circulating pancreas epithelial cells in patients with pancreatic cystic lesions. Gastroenterology 2014;146:647-51. [Crossref] [PubMed]
- Rhim AD, Mirek ET, Aiello NM, et al. EMT and dissemination precede pancreatic tumor formation. Cell 2012;148:349-61. [Crossref] [PubMed]
- Riva F, Dronov OI, Khomenko DI, et al. Clinical applications of circulating tumor DNA and circulating tumor cells in pancreatic cancer. Mol Oncol 2016;10:481-93. [Crossref] [PubMed]
- Trajkovic K, Hsu C, Chiantia S, et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 2008;319:1244-7. [Crossref] [PubMed]
- Skog J, Würdinger T, van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 2008;10:1470-6. [Crossref] [PubMed]
- Kahlert C, Melo SA, Protopopov A, et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem 2014;289:3869-75. [Crossref] [PubMed]
- Melo SA, Luecke LB, Kahlert C, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015;523:177-82. [Crossref] [PubMed]
- Ding G, Zhou L, Qian Y, et al. Pancreatic cancer-derived exosomes transfer miRNAs to dendritic cells and inhibit RFXAP expression via miR-212-3p. Oncotarget 2015;6:29877-88. [PubMed]
- Bauden M, Pamart D, Ansari D, et al. Circulating nucleosomes as epigenetic biomarkers in pancreatic cancer. Clin Epigenetics 2015;7:106. [Crossref] [PubMed]
- Wang S, Chen X, Tang M. Quantitative assessment of the diagnostic role of MUC1 in pancreatic ductal adenocarcinoma. Tumour Biol 2014;35:9101-9. [Crossref] [PubMed]
- Singh N, Gupta S, Pandey RM, et al. High levels of cell-free circulating nucleic acids in pancreatic cancer are associated with vascular encasement, metastasis and poor survival. Cancer Invest 2015;33:78-85. [Crossref] [PubMed]
- Kulemann B, Pitman MB, Liss AS, et al. Circulating tumor cells found in patients with localized and advanced pancreatic cancer. Pancreas 2015;44:547-50. [Crossref] [PubMed]
- Herreros-Villanueva M, Bujanda L, Billadeau DD, et al. Embryonic stem cell factors and pancreatic cancer. World J Gastroenterol 2014;20:2247-54. [Crossref] [PubMed]
- Chari ST, Kelly K, Hollingsworth MA, et al. Early detection of sporadic pancreatic cancer: summative review. Pancreas 2015;44:693-712. [Crossref] [PubMed]