
Genetic changes in rat proximal nerve stumps after sciatic nerve transection
Introduction
Peripheral nerves are unprotected, delicate, and easily broken tissues. Injury to peripheral nerves, especially injury with long nerve gaps, often induces motor, sensory, and autonomic nerve function disorders and leads to long-term disabilities (1,2). Lucky, different from nerves in the adult mammalian central nervous system, nerves in the peripheral nervous system are capable of some level of self-regeneration, although their self-regeneration are limited after serious nerve injury (3,4). Gaining a better understanding of the biological processes of peripheral nerve regeneration may help the discovery of potential therapies for promoting peripheral nerve regeneration after injury and may provide some cues for central nerve regeneration.
Morphological studies demonstrate that two distinct biological processes occur in the proximal nerve stumps and the distal nerve stumps after peripheral nerve injury (5). The distal nerve stumps undergo a degeneration process called Wallerian degeneration within 24h to 36h after peripheral nerve injury (6,7). Macrophages and Schwann cells, the unique glial cells in the peripheral nervous system, serve to remove axon and myelin debris and clear a regenerative path (8-11). The proximal nerve stumps, on the other hand, undergo a regeneration process. During the regeneration process, Schwann cells dedifferentiate, proliferate, and migrate to form a guidance tunnel known as Band of Büngner (12). Moreover, Schwann cells secret cytokines, growth factors, and neurotrophic factors and construct a beneficial microenvironment for axonal elongation and subsequent nerve reinnervation (13,14).
The regeneration process in the proximal nerve stumps is critical for the growth of axons towards the distal nerve stumps and the functional recovery of injured nerves. Therefore, it is essential to investigate the molecular changes in the proximal nerve stumps after peripheral nerve injury to identify critical cues for nerve regeneration. In view of this, we achieved a previous conducted microarray dataset (Gene Expression Omnibus (GEO) database GSE30165) that measured gene expressions in the proximal nerve stumps at 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d after rat sciatic nerve transection. Microarray data was re-analyzed by using Ingenuity pathway analysis (IPA) software (http://www.ingenuity.com/), a commercial software package that commonly used to interpreted high-throughput data (15,16).
Significantly involved canonical signaling pathways, diseases biological processes, and genetic networks in differentially expressed genes in the proximal nerve stumps after nerve transection were revealed. IPA disease and biological function “Organismal Injury and Abnormalities” was investigated in detail and the expression levels of representing genes in “Organismal Injury and Abnormalities”, including S100 Calcium Binding Protein A8 (S100A8), Sulfiredoxin 1 (SRXN1), C-C Motif Chemokine Ligand 20 (CCL20), and Cytochrome P450 Family 1 Subfamily A Member 1 (CYP1A1), were validated.
Methods
Ethics statement
Animals were provided by the Experimental Animal Center of Nantong University, Jiangsu, China. All the experimental procedures involving animals were conducted in accordance with Institutional Animal Care guidelines of Nantong University and approved ethically by the Administration Committee of Experimental Animals, Jiangsu, China.
Animal surgery
Male, adults Sprague-Dawley rat weighting 180–220 g were randomly divided into 10 groups: 0 h, 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d to undergo sciatic nerve transection. Animal surgery was conducted as previously reported (17). Briefly, each rat was first anaesthetized by an intraperitoneal injection of complex narcotics containing 85 mg/kg trichloroacetaldehyde monohydrate (RichJoint, Shanghai, China), 42 mg/kg magnesium sulfate (Xilong Scientific, Guangzhou, Guangdong, China), 17 mg/kg sodium pentobarbital (Sigma-Aldrich, St. Louis, MO, USA). Anaesthetized rat was subjected to a 30 mm-long incision on the lateral side of the mid-thigh of the left hind limb and a 10 mm-long nerve stump excision at the site just proximal to the division of tibial and common peroneal nerves. Rats in the 0 h group received sham-surgery and were used as normal controls. At 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, or 14 d after nerve transection, a 5 mm-long nerve stump at the proximal site was excised, collected, and subjected to microarray analysis and validation experiments.
Microarray analysis
Total RNAs were extracted from rat proximal nerve stumps using Trizol (Invitrogen, Carlsbad, CA, USA). The quantitation and quality of extracted RNAs were assessed using a Nanodrop ND-1000 spectrophotometer (Infinigen Biotechnology Inc., City of Industry, CA, USA) and a BioAnalyzer 2100 (Agilent Technology, Santa Clara, CA, USA), respectively. Microarray analysis was carried out using the Affymetrix GeneChip Hybridization Oven 640, Gene Array Scanner 3000 (Affymetrix, Santa Clara, CA, USA), and GeneChip Scan Control software (Affymetrix). Microarray data were submitted to a online GEO database (GSE30165).
Bioinformatic analysis
The expression levels of genes at each time point (0 h, 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d) after nerve transection in the proximal nerve stumps were determined. Gene expression levels were up-loaded to IPA software (Ingenuity Systems Inc., Redwood City, CA, USA). Genes with an experimental log ratio 1 and an adjusted P value <0.05 as compared with 0 h control were considered as differentially expressed and were conducted for IPA core analysis. Differentially expressed genes in the proximal nerve stumps were overlaid with the Ingenuity pathway knowledge base (IPKB) to achieve significantly enriched canonical signaling pathways, diseases and biological functions, and genetic networks.
Quantitative real-time PCR
Quantitative real-time PCR was performed by using RNA samples different from those used for microarray. cDNA was achieved from extracted RNAs using the Prime-Script reagent Kit (TaKaRa,Dalian,China) and then used as a template for gene amplification using SYBR Green Premix Ex Taq (TaKaRa) using an Applied Biosystems Stepone real-time PCR System. The sequences of primers pairs were follows: S100A8 (forward) 5'-GGGAATCACCATGCCCTCTA-3' and (reverse) 5'-CCCACCCTTATCACCAACACA-3'; SRXN1 (forward) 5'-ACCTCCTGATACCCCACTCC-3' and (reverse) 5'-GGAACCCCCTCATTCTTGGG-3'; CCL20 (forward) 5'-AGTCAGAAGCAGCAAGCAACTTT-3‘ and (reverse) 5'-CTTCGTCGGCCATCTGTGTT-3'; CYP1A1 (forward) 5'-CCTGGGGTCCTAGAGAACACT-3' and (reverse) 5'-AACCACCCAGAATCCAAGGC-3'; and GAPDH (forward) 5'-ACAGCAACAGGGTGGTGGAC-3' and (reverse) 5'-TTTGAGGGTGCAGCGAACTT-3'. The thermocycler program was as follows: 5 min at 95 °C; 40 cycles of 30 s at 95 °C, 45 sec at TAnneal, and 30 s at 72 °C; and 5 min at 72 °C. Relative expression levels of target genes S100A8, SRXN1, CCL20, and CYP1A1 were determined using the comparative 2−ΔΔCt method using GAPDH as the internal control.
Statistical analysis
Data were presented as means ± SEM. Multiple comparisons between nerve transected rats (0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d) and uninjured rats (0 h) were performed using one-way ANOVA and Dunnett’s test. A P value <0.05 was considered as statistically significant.
Results
Significantly enriched canonical signaling pathways in the proximal nerve stumps after rat nerve transection
A total of 41,092 gene IDs were up-loaded to IPA software and a total of 16,269 gene IDs were mapped to the IPKB database. Differentially expressed genes with an experimental log ratio 1 and an adjusted P value <0.05 were screened from these mapped gene IDs and categorized to IPA canonical signaling pathways (http://fp.amegroups.cn/cms/f99f5422c71ba457cdb59f3e839e9614/atm.2019.11.98-1.pdf). IPA canonical signaling pathways with a P value <10−5 (1.00E-05) at least one time point (0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, or 14 d) after sciatic nerve transection were listed and their corresponding P values were labeled (Figure 1). These enriched IPA canonical signaling pathways were ranked with increased P values from 0.5 h to 14 d and displayed in color gradation with red color indicated most significantly enriched canonical signaling pathway. Non significantly enriched canonical signaling pathways at some time points were labeled in green color and labeled with a P value >0.05 (>5.00E-02).

Only one single canonical signaling pathway, “Airway Pathology in Chronic Obstructive Pulmonary Disease”, was enriched in the proximal nerve stumps at the very early time point (0.5 h) after rat sciatic nerve transection. A larger number of canonical signaling pathways were significantly activated at later time points (1, 3, 6, and 9 h). Then, the number of significantly activated canonical signaling pathways seemed to decrease. These canonical signaling pathways could mainly be categorized to cytokine signaling (“Airway Pathology in Chronic Obstructive Pulmonary Disease” and “Role of IL-17A in Psoriasis”), cellular immune response (“Agranulocyte Adhesion and Diapedesis”, “Granulocyte Adhesion and Diapedesis”, “Role of Cytokines in Mediating Communication between Immune Cells”, “IL-10 Signaling”, “IL-6 Signaling”, “Graft-versus-Host Disease Signaling”, and “Communication between Innate and Adaptive Immune Cells”), nuclear receptor signaling (“LXR/RXR Activation” and “LPS/IL-1 Mediated Inhibition of RXR Function”), disease-specific pathways (“Hepatic Cholestasis”, “Atherosclerosis Signaling”, “Role of Macrophages, Fibroblasts and Endothelial Cells in Rheumatoid Arthritis”, and “Role of Hypercytokinemia/hyperchemokinemia in the Pathogenesis of Influenza”), and organismal growth and development (“Inhibition of Matrix Metalloproteases” and “Hematopoiesis from Pluripotent Stem Cells”).
Significantly enriched diseases and biological functions in the proximal nerve stumps after rat nerve transection
Besides significantly involved canonical signaling pathways, significantly involved diseases and biological functions in the proximal nerve stumps after nerve transection were also revealed (http://fp.amegroups.cn/cms/e54e63493fea7caf27d8fe939ab12f20/atm.2019.11.98-2.pdf). IPA diseases and biological functions with a P value <10−10 (1.00E-10) at least one time point (0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, or 14 d) after nerve transection were listed, ranked, and marked in colors (Figure 2). Consistent with previous observation of significantly involved canonical signaling pathways, few diseases and biological functions were found to be highly involved at early time points, numerous diseases and biological functions were involved at later time points, and less diseases and biological functions were involved in even longer period of time after nerve transection.

Significantly involved diseases and biological functions were mainly grouped to organ development (“Hematological System Development and Function”, “Tissue Morphology”, “Cardiovascular System Development and Function”, and “Organismal Development”), inflammation and immune response (“Inflammatory Response”, “Immune Cell Trafficking”, and “Inflammatory Disease”), diseases and organ abnormalities (“Organismal Injury and Abnormalities”, “Hematological Disease”, “Immunological Disease”, and “Dermatological Diseases and Conditions”), as well as cellular behavior (“Cellular Movement” and “Cell-To-Cell Signaling and Interaction”).
Diseases and biological functions “Organisamal Injury and Abnormalities”
IPA disease and biological function “Organismal Injury and Abnormalities” was subjected to further study considering that this disease and biological function term was directly related with nerve injury. Differentially expressed genes in “Organismal Injury and Abnormalities” were listed in http://cdn.amegroups.cn/static/application/96781c4b87047c21a78f984f0e74760b/atm.2019.11.98-3.pdf. The expression patterns of these differentially expressed genes in the proximal nerve stumps at 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d after nerve transection were displayed in a heatmap (Figure 3).
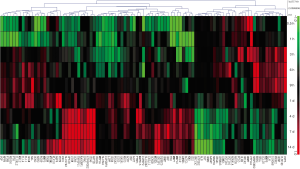
The heatmap showed that about 1/4 of differentially expressed genes were down-regulated in the proximal nerve stumps at 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d after nerve transection. About 1/3 of differentially expressed genes were first up-regulated and then down-regulated and the majority of differentially expressed genes were up-regulated after nerve transection (Figure 3).
Representative genes were selected for quantitative real-time PCR validation. The expression levels of S100A8, SRXN1, CCL20, and CYP1A1 in the proximal nerve stumps at all tested time points were measured and normalized to the internal control GAPDH. Outcomes from quantitative real-time PCR were in consistent with outcomes from microarray analysis, in that S100A8, SRXN1, and CCL20 were first up-regulated and then down-regulated in the proximal nerve stumps after nerve transection and CYP1A1 was kept down-regulated after nerve transection (Figure 4).
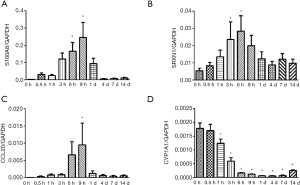
Genetic networks
The interactions and networks between differentially expressed genes in various biological functions were then analyzed by IPA genetic networks. IPA genetic networks of differentially expressed genes in the proximal nerve stumps at 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, or 14 d after nerve transection were scored, ranked, and listed in http://fp.amegroups.cn/cms/85b0b2c8314b2869c97df493626ef67c/atm.2019.11.98-4.pdf. Top 3 IPA genetic networks at each time point were screened and listed in Table 1. The top enriched IPA genetic network in Table 1 obtained a rating score of 30, suggesting that the possibility that genes in these networks were not connected was less than 10−30. Even the lowest score in Table 1 reached 17, indicating that these genes were highly interacted with each other and these biological functions were closely related.

Full table
Notably, IPA disease and biological function “Organismal Injury and Abnormalities” were present in IPA genetic networks at 0.5 h (network #3), 9 h (network #3), and 1 d (network #1) after nerve transection. These “Organismal Injury and Abnormalities”—involved genetic networks at 0.5 h, 9 h, and 1d were displayed to demonstrate gene interactions and hub genes (Figure 5). Genetic networks showed that Sodium Voltage-Gated Channel Beta Subunit 2 (SCN2B) was the hub gene at 0.5 h after nerve transection and the expression of SCN2B was down-regulated at this time point (Figure 5A). Down-regulated CYP1A1 and up-regulated S100A8 and PR Domain Containing 1, With ZNF Domain (PRDM1) seemed to be hub genes at 9 h after nerve transection (Figure 5B) while down-regulated CYP1A1 and up-regulated C-Reactive Protein (CRP) and Interleukin 11 (IL11) seemed to be hub genes at 1d after nerve transection (Figure 5C).
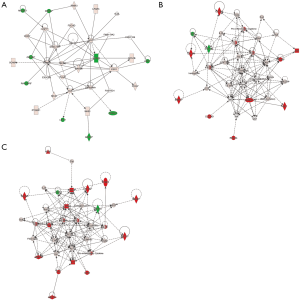
Discussion
Recently, with the development of high-throughput analysis tools such as DNA microarray, protein array, DNA sequencing, and RNA sequencing, molecular changes following peripheral nerve injury were also been largely revealed. For example, RNA deep sequencing previously conducted in our laboratory identified numerous differentially expressed RNAs as well as alternatively spliced genes at the injury sites in rat sciatic nerve stumps at 1, 4, 7, and 14 days after nerve crush (18,19). This RNA deep sequencing, however, analyzed the entire injure sites and did not separate the proximal and the distal nerve stumps. To clearly demonstrate molecular changes during Wallerian degeneration, gene and protein expression profiling in the distal nerve stumps after rat nerve injury were investigated and critical biological activities were discovered (20-24). Moreover, gene expression patterns in the proximal nerve stumps and the distal nerve stumps after rat sciatic nerve injury were jointly investigated and the expression levels of some genes in different localizations were compared (25). Still, as far as we know, there exists no systemical analysis of genetic changes in the proximal nerve stumps at multiple time points after peripheral nerve injury. Conducting relevant study will fill the knowledge gap and lead to a deeper understanding of nerve regeneration process.
In the current study, by investigating previously obtained microarray dataset using an advanced bioinformatic tool, we screened differentially expressed genes in the proximal nerve stumps at 0.5 h, 1 h, 3 h, 6 h, 9 h, 1 d, 4 d, 7 d, and 14 d after rat sciatic nerve transection and discovered signaling enriched signaling pathways and biological functions at each time point. IPA study revealed that lots of signaling pathways and biological functions were activated in the proximal nerve stumps at 3, 6, and 9 h after nerve transection. These activated signaling pathways and biological functions were mainly associated with cellular and organ development, inflammation and immune response, and diseases and organ abnormalities. Consistent with previous bioinformatic analysis of differentially expressed genes in the sciatic nerve crush sites and distal nerve stumps (21,26), some inflammation and immune response-related canonical signaling pathways, such as “Agranulocyte Adhesion and Diapedesis”, “Granulocyte Adhesion and Diapedesis”, “IL-10 Signaling”, and “IL-6 Signaling”, were activated from as early as 1h after nerve injury. This indicated that inflammation and immune cascades, as important protective response and feedbacks against neural injury (27,28), were not only essential for debris clearance in the distal nerve stumps during Wallerian degeneration, but also critical for nerve regeneration in the proximal nerve stumps.
Detailed analysis of IPA disease and biological function “Organismal Injury and Abnormalities” showed that numerous genes in this biological function were differentially expressed. Subsequent validation with quantitative real-time PCR demonstrated that S100A8, SRXN1, and CCL20 were up-regulated in the proximal nerve stumps and CYP1A1 was down-regulated in the proximal nerve stumps after nerve transection. S100A8, a gene encoding for an acidic calcium-binding protein, is correlated with various inflammatory conditions by inducing cytokine secretion and immune cell infiltration (29). Notably, S100A8 was found to be one of the top up-regulated genes in both the proximal and distal nerve stumps at 1 day after injury (30). Our current study demonstrated that the expressions of S100A8 in proximal nerve stumps was elevated at even earlier time points, indicating the early involvement of S100A8 in nerve regeneration. Besides S100A8, CCL20 is also associated with inflammation and immune response. For example, CCL20 has been demonstrated to be highly expressed in experimental autoimmune encephalomyelitis, a inflammatory disease of the central nervous system (31). Here, the high abundances of CCL20 further demonstrated the significant involvement of inflammation and immune response in the regeneration process. SRXN1 and CYP1A1, two other validated genes, have not been commonly reported in peripheral nerve repair and regeneration. Therefore, their biological roles need to be further determined.
Conclusions
Taken together, our current study, from the aspect of transcriptome analysis, identified comprehensive changes of genes in the proximal nerve stumps after rat sciatic nerve transection injury. These robust genetic changes may be critical for the regeneration process and may contribute to the discovery of new therapeutic methods for peripheral nerve injury.
Acknowledgments
Funding: This work was supported by the National Natural Science Foundation of China (81971170), the health committee project of Jiangsu province (H2018023), the health committee major project of Wuxi City (Z201702).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. Animals were provided by the Experimental Animal Center of Nantong University, Jiangsu, China. All the experimental procedures involving animals were conducted in accordance with Institutional Animal Care guidelines of Nantong University and approved ethically by the Administration Committee of Experimental Animals, Jiangsu, China.
References
- Yi S, Xu L, Gu X, et al. Scaffolds for peripheral nerve repair and reconstruction. Exp Neurol 2019;319:112761. [Crossref] [PubMed]
- Gu X, Ding F, Yang Y, et al. Construction of tissue engineered nerve grafts and their application in peripheral nerve regeneration. Prog Neurobiol 2011;93:204-230. [Crossref] [PubMed]
- Chen ZL, Yu WM, Strickland S. Peripheral regeneration. Annu Rev Neurosci 2007;30:209-33. [Crossref] [PubMed]
- Raimondo S, Fornaro M, Tos P, et al. Perspectives in regeneration and tissue engineering of peripheral nerves. Ann Anat 2011;193:334-40. [Crossref] [PubMed]
- Geuna S, Raimondo S, Ronchi G, et al. Chapter 3: Histology of the peripheral nerve and changes occurring during nerve regeneration. Int Rev Neurobiol 2009;87:27-46. [Crossref] [PubMed]
- Conforti L, Gilley J, Coleman MP. Wallerian degeneration: an emerging axon death pathway linking injury and disease. Nat Rev Neurosci 2014;15:394-409. [Crossref] [PubMed]
- Eder M, Schulte-Mattler W, Poschl P. Neurographic course Of Wallerian degeneration after human peripheral nerve injury. Muscle Nerve 2017;56:247-52. [Crossref] [PubMed]
- Coleman MP, Conforti L, Buckmaster EA, et al. An 85-kb tandem triplication in the slow Wallerian degeneration (Wlds) mouse. Proc Natl Acad Sci U S A 1998;95:9985-90. [Crossref] [PubMed]
- Chen P, Piao X, Bonaldo P. Role of macrophages in Wallerian degeneration and axonal regeneration after peripheral nerve injury. Acta Neuropathol 2015;130:605-18. [Crossref] [PubMed]
- Gaudet AD, Popovich PG, Ramer MS. Wallerian degeneration: gaining perspective on inflammatory events after peripheral nerve injury. J Neuroinflammation 2011;8:110-22. [Crossref] [PubMed]
- Wong KM, Babetto E, Beirowski B. Axon degeneration: make the Schwann cell great again. Neural Regen Res 2017;12:518-24. [Crossref] [PubMed]
- Bhatheja K, Field J. Schwann cells: origins and role in axonal maintenance and regeneration. Int J Biochem Cell Biol 2006;38:1995-9. [Crossref] [PubMed]
- Clements MP, Byrne E, Camarillo Guerrero LF, et al. The Wound Microenvironment Reprograms Schwann Cells to Invasive Mesenchymal-like Cells to Drive Peripheral Nerve Regeneration. Neuron 2017;96:98-114.e7. [Crossref] [PubMed]
- Jessen KR, Mirsky R, Lloyd AC. Schwann Cells: Development and Role in Nerve Repair. Cold Spring Harb Perspect Biol 2015;7:a020487. [Crossref] [PubMed]
- Jiménez-Marín Á, Collado-Romero M, Ramirez-Boo M, et al. Biological pathway analysis by ArrayUnlock and Ingenuity Pathway Analysis. BMC Proc 2009;3 Suppl 4:S6. [Crossref] [PubMed]
- Krämer A, Green J, Pollard J Jr, et al. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 2014;30:523-30. [Crossref] [PubMed]
- Yu B, Zhou S, Wang Y, et al. Profile of microRNAs following rat sciatic nerve injury by deep sequencing: implication for mechanisms of nerve regeneration. PLoS One 2011;6:e24612. [Crossref] [PubMed]
- Yi S, Zhang H, Gong L, et al. Deep Sequencing and Bioinformatic Analysis of Lesioned Sciatic Nerves after Crush Injury. PLoS One 2015;10:e0143491. [Crossref] [PubMed]
- Zhao L, Yi S. Transcriptional landscape of alternative splicing during peripheral nerve injury. J Cell Physiol 2019;234:6876-85. [Crossref] [PubMed]
- Yi S, Tang X, Yu J, et al. Microarray and qPCR Analyses of Wallerian Degeneration in Rat Sciatic Nerves. Front Cell Neurosci 2017;11:22. [Crossref] [PubMed]
- Yu J, Gu X, Yi S. Ingenuity Pathway Analysis of Gene Expression Profiles in Distal Nerve Stump following Nerve Injury: Insights into Wallerian Degeneration. Front Cell Neurosci 2016;10:274. [Crossref] [PubMed]
- Li M, Zhang P, Guo W, et al. Protein expression profiling during wallerian degeneration after rat sciatic nerve injury. Muscle Nerve 2014;50:73-8. [Crossref] [PubMed]
- Yao D, Li M, Shen D, et al. Gene expression profiling of the rat sciatic nerve in early Wallerian degeneration after injury. Neural Regen Res 2012;7:1285-92. [PubMed]
- Li M, Guo W, Zhang P, et al. Signal flow and pathways in response to early Wallerian degeneration after rat sciatic nerve injury. Neurosci Lett 2013;536:56-63. [Crossref] [PubMed]
- Jiang N, Li H, Sun Y, et al. Differential gene expression in proximal and distal nerve segments of rats with sciatic nerve injury during Wallerian degeneration. Neural Regen Res 2014;9:1186-94. [Crossref] [PubMed]
- Xing L, Cheng Q, Zha G, et al. Transcriptional Profiling at High Temporal Resolution Reveals Robust Immune/Inflammatory Responses during Rat Sciatic Nerve Recovery. Mediators Inflamm 2017;2017:3827841. [PubMed]
- Benowitz LI, Popovich PG. Inflammation and axon regeneration. Curr Opin Neurol 2011;24:577-83. [Crossref] [PubMed]
- Donnelly DJ, Popovich PG. Inflammation and its role in neuroprotection, axonal regeneration and functional recovery after spinal cord injury. Exp Neurol 2008;209:378-88. [Crossref] [PubMed]
- Simard JC, Cesaro A, Chapeton-Montes J, et al. S100A8 and S100A9 induce cytokine expression and regulate the NLRP3 inflammasome via ROS-dependent activation of NF-kappaB(1.). PLoS One 2013;8:e72138. [Crossref] [PubMed]
- Chernov AV, Dolkas J, Hoang K, et al. The calcium-binding proteins S100A8 and S100A9 initiate the early inflammatory program in injured peripheral nerves. J Biol Chem 2015;290:11771-84. [Crossref] [PubMed]
- Evangelista MG, Castro SB, Alves CC, et al. Early IFN-gamma production together with decreased expression of TLR3 and TLR9 characterizes EAE development conditional on the presence of myelin. Autoimmunity 2016;49:258-67. [Crossref] [PubMed]


