
Engineered exosomes for targeted delivery of miR-187-3p suppress the viability of hemangioma stem cells by targeting Notch signaling
Introduction
Infantile hemangioma (IH) is the most common benign vascular tumor of infancy, affecting approximately 10% of infants. It is more likely to occur in White, female, and premature infants (1). IH is proposed to arise from hemangioma stem cells (HemSCs) during infancy and has a unique ability to proliferate rapidly (2). Unfortunately, a subset of patients with IH develop complications, including pain, functional impairment, and permanent visual impairment, which can lead to significant morbidity and mortality. Common therapies for IH include oral beta-blockers, surgery, and laser therapy (2). For patients with IH, early intervention during the proliferative phase can minimize the risk of developing potential complications. Therefore, it is imperative to understand the mechanisms responsible for rapid IH cell proliferation in order to treat and prevent the disease.
Exosomes are nano-sized extracellular vesicles that are secreted by most cell types, including stem cells and cancer cells (3). Numerous studies have demonstrated exosomes to play a powerful role in the mediation of cell-cell communication via the delivery of effectors, such as proteins, messenger RNAs, or microRNAs (miRNAs), to target cells. These natural nanovesicles can transmit crucial cellular information and can be engineered to have robust delivery and targeting capacity. Mesenchymal stem cells (MSCs) have been widely used to produce exosomes due to their accessibility and availability from ethically acceptable sources, such as fat tissue (4).
Over the past decade, miRNAs have been one of the best-studied non-coding RNAs and play a critical role in various biological and pathological processes (5,6). Some miRNAs have been reported to be aberrantly expressed in IH. For example, Strub et al. (7) identified chromosome 19 miRNA cluster (C19MC) as the first circulating biomarker of IH, reporting that levels of circulating C19MC were correlated with IH tumor size and propranolol treatment response. Mong et al. (8) discovered the role of the LIN28B/let-7 switch in the pathogenesis of IH and provided a novel mechanism by which propranolol induces IH involution. Zeng et al. (9) found that downregulation of miR-501 inhibited hemangioma cell proliferation, cell cycle progression, colony formation, migration, and invasion in vitro.
miR-187-3p is a novel cancer-related miRNA that has been confirmed to promote or inhibit various malignancies (10,11). However, its role in the development and progression of IH remains unclear. Previously, it was determined that human adipose MSC-derived exosomes (hAMSC-exos), with outstanding biocompatibility and easy accessibility, could serve as a nanoplatforms for delivering miRNAs and improve the bioavailability of exogenous miRNAs (12).
The present study exploited engineered exosomes (E-exos) to deliver miR-187-3p into HemSCs. The E-exos were generated by introducing miR-187-3p mimics into hAMSC-exos via electroporation. We hypothesized that miR-187-3p would inhibit the viability and proliferation of HemSCs by targeting Notch signaling and that using hAMSC-exos as the delivery carrier would greatly facilitate this process, possibly enhancing the physiological effects relative to those of free miR-187-3p. To verify this hypothesis, we investigated the effects of miR-187-3p-containing E-exos on HemSC viability and proliferation in vitro. This study provides a foundation for the future use of hAMSC-exos to deliver drugs and to optimize current clinical options for the treatment of IH. We present the following article in accordance with the MDAR reporting checklist (available at https://atm.amegroups.com/article/view/10.21037/atm-21-4138/rc).
Methods
Isolation of hAMSC-exos from human adipose MSCs
Human adipose MSCs (hAMSCs; HUXMD-01001; P2) were purchased from Cyagen Biotechnology (Cyagen Biotechnology, Guangzhou, China). The hAMSCs were cultured in Human Adipose-derived Mesenchymal Stem Cell Basal Medium (HUXMD-01001; Cyagen Biotechnology) containing 10% fetal bovine serum (FBS; A4766801, Gibco, Waltham, MA, USA), 1% penicillin/streptomycin (10378016; Thermo Scientific, Waltham, MA, USA), and 1% glutamine (1294808; Sigma-Aldrich, Darmstadt, Germany). When approximately 60–70% confluence was reached, the culture medium was discarded and the hAMSCs were rinsed three times with phosphate-buffered saline (PBS; E607008-0500; Sangon Biotech, Shanghai, China). Then, the hAMSCs were cultured in conditioned medium supplemented with exosome-depleted FBS (EXO-FBS-250A-1; System Biosciences, Mountain View, CA, USA) for subsequent exosome harvesting. Exosomes were isolated by ultracentrifugation according to methods reported previously (13). Briefly, after 48-hour culture, the hAMSC conditioned medium was collected. Dead cells and cell debris were removed by centrifugation (4 ℃, 300 ×g for 10 minutes). The supernatant was collected and filtered through a 0.22-µm filter (GSWP04700; Merck Millipore, Darmstadt, Germany ) and then ultracentrifuged at 100,000 ×g for 6 hours. The resulting supernatant was discarded, and the pellet containing the exosomes was retained. All procedures for isolating exosomes from the conditioned medium were performed at 4 ℃.
Isolation and culture of HemSCs
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Institute Review Board of Shanghai Ninth People’s Hospital (No. SH9H-2019-T272-1) and informed consent was taken from the parents or guardians of all the patients.
Specimens of proliferating IH were collected from the IH patients visiting the Department of Oromaxillofacial Head and Neck Oncology, Shanghai Ninth People’s Hospital between August 2016 and October 2018. HemSCs were isolated from specimens of proliferating IH as described previously (14). They were cultured in endothelial basal medium (EBM-2; CC-3156; Lonza, Basel, Switzerland) supplemented with 20% FBS (A4766801; Gibco) at 37 ℃ in a 5% CO2 humidified atmosphere.
Transmission electron microscopy (TEM) and dynamic light scattering (DLS)
To determine whether the isolated extracellular vesicles were exosomes, the morphology and size distribution of the isolated particles were characterized by TEM and DLS. The morphology of hAMSC-exos was evaluated by TEM as described previously (15). Briefly, exosomes were resuspended in PBS and then the suspension was dropped on a carbon-coated copper grid, stained with 2% uranyl acetate, and imaged under a Hitachi H7500 TEM microscope (Tokyo, Japan) at 80 kV. The size distribution and exosome concentration in each group were determined using a NanoSight LM10 (Malvern Instruments, Malvern, UK), as described previously (16). All samples were diluted with PBS to a concentration of approximately (1–5)×1010 particles/mL. The hydrodynamic sizes of the purified exosomes were measured by DLS. DLS measurements were conducted at 25 ℃ on a Zetasizer Nano ZS (Malvern Instruments). The concentration of exosome pellets was 10 mg/mL.
Western blot analysis and quantitative reverse transcription-polymerase chain reaction (qRT-PCR)
Western blot analysis was performed as described previously (17). Primary antibodies against heat shock protein 70 (Hsp70; 1:1,000 dilution; ab2787) and tumor susceptibility gene 101 (TSG101; 1:1,000 dilution; ab125011) were purchased from Abcam (Cambridge, UK), and anti-CD9 antibodies (1:1,000 dilution; #13174) were purchased from Cell Signaling Technology (Danvers, MA, USA). Samples were incubated with each primary antibody in turn at 4 ℃ overnight, and specific proteins were imaged using a FluorChem Q system (ProteinSimple, San Jose, CA, USA).
For qRT-PCR analysis, total exosomal miRNAs were extracted using TRIzol reagent (15596026; Invitrogen, Waltham, MA, USA). The expression levels of miR-187-3p were detected using a SYBR green (4364346; Applied Biosystems, Waltham, MA, USA) assay according to the manufacturer’s instructions. U6 expression was used as control for miR-187-3p. Relative expression was analyzed using the 2−ΔΔCT method. All samples were analyzed in triplicate.
The primers were used for qRT-PCR were as follows: miR-187-3p, 5'-CTT CGT GTC TTG TGT TGC-3' (forward) and 5'-GTG CAG GGT CCG AGG T-3' (reverse); U6, 5'-TGC GGG TGC TCG CTT CGG CAG C-3' (forward) and 5'-GTG CAG GGT CCG AGG T-3' (reverse).
Preparing E-exos by electroporation
To load the exosomes with miR-187-3p, E-exos were prepared based on the methods of Alvarez-Erviti et al. (18). The miR-187-3p mimics, inhibitor, and control were purchased from Genomeditech (Shanghai, China). First, hAMSC-exos were resuspended with Bovine Serum Albumin (23209; Thermo Scientific) to a total protein concentration of 12 µg/mL [measured using a BCA kit (C503061; Sangon Biotech)]. Then, 200 nM miR-187-3p mimics or miR-187 negative control (NC) was mixed with 50 µL electroporation buffer (MPK10025; Neon transfection kit; Invitrogen) at 4 ℃. Following that, 3 mL prechilled electrolytic buffer (MPK10025; Neon transfection kit; Invitrogen) was added to the Neon tube. The mixture was aspirated into the Neon pipette, which was inserted into the Neon tube. The whole tube was inserted into the Neon Pipette Station (MPK500; Invitrogen). Then, electroporation was performed once at 500 V with a 1-ms pulse duration. After the pulse, the mixture was transferred to a new tube and incubated for 30 minutes at 37 ℃. The free miR-187-3p mimics were removed through ultracentrifugation (657,000 ×g, 6 hours, 4 ℃). The final engineered exosomes (E-exos loaded with miR-187-3p mimics or the NC) were isolated again by ultracentrifugation (100,000 ×g, 60 minutes, 4 ℃), suspended in PBS, and stored at −80 ℃ until use. The sequences of the miR-187 mimics were 5'-UCGUGUCUUGUGUUGCAGCCGG-3' and 3'-UUAGCACAGAACACAACGUCGG-5', whereas the sequences of the miR-187 NC were 5'-UUCUCCGAACGUGUCACGUTT-3' and 3'-TTAAGAGGCUUGCACAGUGCA-5'.
HemSC uptake of E-exos
Labeling of hAMSC-exos and E-exos was performed using the CellTracker CM-Dil kit (180854-97-1; Invitrogen). Briefly, exosomes were resuspended in PBS and incubated with 5 µM CM-Dil dye at 37 ℃ for 15 minutes in the dark. The free CM-Dil dye was subsequently removed via ultracentrifugation at 100,000 ×g for 15 minutes at 4 ℃. The labeled exosomes were rinsed twice with PBS and stored at 4 ℃. HemSCs were incubated with 5 µg/mL Dil-labeled exosomes for 4 hours at 4 ℃. After the supernatants had been discarded, samples were washed with PBS, fixed with 4% paraformaldehyde at 37 ℃ for 15 minutes, washed three times with PBS, and finally incubated with Hoechst (C1017; Beyotime Biotechnology, Shanghai, China) at 37 ℃ for 5 minutes. Samples were then imaged using a confocal laser scanning microscope (FV3000; Olympus, Tokyo, Japan).
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay
The effects of E-exos on the viability and proliferation of HemSCs were evaluated using an MTT assay. HemSCs were seeded into 96-well plates at density of 1×104 cells/well and then cultured for 8 hours at 37 ℃ in a 5% CO2 atmosphere. Then, cells were treated with PBS, free miR-187-3p mimics, hAMSC-exos, E-exos (NC), or E-exos for an additional 24 hours. The supernatants were then removed, and 180 µL fresh medium and 20 µL of 5 mg/mL MTT reagent (ST1537-1g; Beyotime Biotechnology) were added to each well. Samples were then incubated at 37 ℃ for 4 hours, the medium was then removed, and the crystals in each well were dissolved in 200 µL dimethylsulfoxide (ST038-100ml; Beyotime Biotechnology). The plates were then incubated at 37 ℃ for 30 minutes before being transferred to a microplate reader (BioTek Synergy HTX). Optical density (OD) was measured at 490 nm. All samples were analyzed in triplicate.
Tube formation assay
HemSCs were cultured on Matrigel-coated plates with EBM-2 supplemented with 20% FBS (A4766801; Gibco) at 37 ℃ in a 5% CO2 humidified atmosphere. When the cells reached 70–80% confluence, HemSC suspensions were prepared by trypsinizing the cells and then resuspending them in culture medium supplemented with 10% FBS (A4766801; Gibco). The HemSCs were treated with either EBM-2 medium (control group), free miR-187-3p mimics, hAMSC-exos, E-exos (NC), or E-exos, cultured for 12 hours at 37 ℃ in a 5% CO2 atmosphere, and then imaged under an inverted microscope (Eclipse Ts2R; Nikon, Tokyo, Japan). Free miR-187-3p mimics, hAMSC-exos, E-exos (NC), and E-exos were administrated at three different concentrations (6, 9, and 12 µg/mL). All samples were analyzed in triplicate.
Statistical analysis
Quantitative data are presented as the mean ± SD. The significance of differences between groups or treatments was measured using Student’s t-test or analysis of variance (ANOVA) followed by the Bonferroni post test. Data were analyzed using GraphPad Prism 8.0. P<0.05 was considered significant. All experiments were repeated in triplicate.
Results
Preparation and identification of E-exos
To determine whether extracellular particles could act as effective carriers for the delivery of miR-187 to target cells, we first isolated hAMSC-exos to investigate the basic characteristics of isolated naïve exosomes. TEM analysis verified that the naturally isolated extracellular particles were dish-shaped vesicles with a double-layer membrane structure (Figure 1A). Next, we used electroporation to load hAMSC-exos with miR-187 NC and miR-187 mimics, to obtain E-exos (NC) and E-exos, respectively. Using TEM (Figure 1B) and DLS analysis (Figure 1C), we found that there were no differences in morphology or size distribution between hAMSC-exos and E-exos; both exhibited a similar size distribution, with a peak diameter of 110 nm (Figure 1C). Finally, western blot analysis revealed that the expression levels of the exosome-specific markers CD9, Hsp70, and TSG101 were similar between hAMSC-exos and E-exos (Figure 1D). These results show that the morphology, particle size, and surface markers of E-exos generated by electroporation did not differ significantly from those of hAMSC-exos.
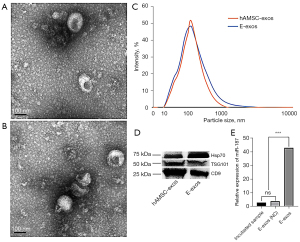
To determine whether miRNAs had been successfully loaded into hAMSC-exos, the hAMSC-exos and the E-exos were lysed separately to detect the control and miR-187-3p mimics. Using qRT-PCR, the relative expression of miR-187-3p in the E-exos group was found to be significantly higher than that in the E-exos (NC) group, and higher than that in the incubated group (i.e., hAMSC-exos incubated with free miR-187 for 30 minutes; Figure 1E). These data suggested that exogenous miR-187-3p had been successfully loaded into hAMSC-exos by electroporation, which indicated that E-exos had been successfully generated and that they met the required characterization criteria of exosomes described in previous studies (19,20).
Effects of miR-187-3p-loaded E-exos on HemSC viability in vitro
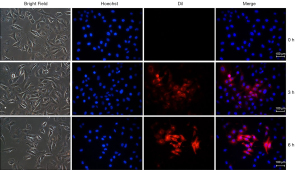
We next incubated HemSCs with E-exos and analyzed their uptake of miR-187-3p. We found that E-exos could be internalized into HemSCs and that the course of cellular E-exos uptake was time dependent. These results indicated that E-exos could serve as effective carriers for the delivery of miR-187 to recipient cells (Figure 2).
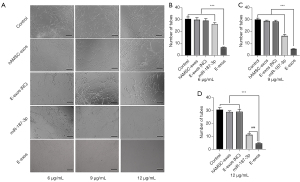
To further explore the potential effects of exogenous miR-187-3p on HemSCs, we used the tube formation assay to evaluate E-exos regulation of the biological behavior of HemSCs. Compared with HemSCs incubated with 6, 9, and 12 µg/mL hAMSC-exos, those incubated with 6, 9, and 12 µg/mL free miR-187-3p were respectively less able to form capillary-like tubular structures after 12-hour culture. Further, the ability of HemSCs to form tube-like structures declined significantly with increasing concentrations of E-exos. As shown in Figure 3, extensive tube networks were observed in HemSCs incubated with EBM-2 medium (control group), hAMSC-exos, and E-exos (NC). In contrast, free miR-187-3p and E-exos significantly inhibited tube formation after 12-hour culture (Figure 3). Moreover, inhibition of tube formation was slightly enhanced with E-exos compared to with free miR-187-3p after 12-hour culture (Figure 3). However, these findings appear inconsistent with the superior effects of E-exos on cell proliferation. We speculate that this could be attributed to the fact that the HemSC tube formation assay was conducted within a short period of time, during which only limited amounts of E-exos were integrated into HemSCs due to the time-dependent uptake.
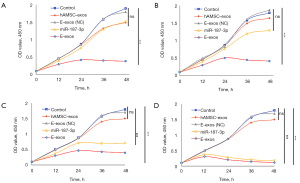
The MTT assay showed that E-exos inhibited HemSCs cell viability at low (6 µg/mL; Figure 4A), medium (9 and 12 µg/mL; Figure 4B,4C), and high (24 µg/mL; Figure 4D) concentrations. There were significant differences in cell viability between all E-exos-treated groups (low, medium, and high concentrations) and the control group (Figure 4), but there was no significant difference between the hAMSC-exos-treated (Figure 4A-4D) or free miR-187-treated (Figure 4C,4D) group and the control group. Together, these findings indicate that E-exos loading with miR-187-3p significantly inhibits the viability of HemSCs in vitro, suggesting that hAMSC-exos are efficient in enhancing miR-187-3p bioavailability.
Role of the Notch pathway in E-exos inhibition of HemSC viability
Next, we evaluated how the viability of HemSCs was affected by E-exos and the potential underlying molecular mechanism. Previous studies have shown that Notch signaling plays an important role in the regulation of HemSC proliferation (21-23). Consistent with the results reported above, E-exos loading with miR-187-3p significantly inhibited the viability of HemSCs in vitro compared with free miR-187 or hAMSC-exos. However, the addition of the Notch signaling inhibitor BMS 299897 (Tocris Bioscience, Minneapolis, MN, USA), which inhibits γ-secretase, reversed this phenotype to a state close to that in the control group. This result indicates that the E-exos-induced decrease in cell viability may be regulated through Notch signaling.
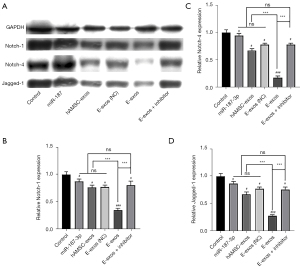
To verify the above hypothesis, HemSCs were treated with free miR-187-3p mimics, hAMSC-exos, E-exos (NC), E-exos, or E-exos plus the γ-secretase inhibitor. Then, proteins related to Notch signaling were detected in each group through western blot analysis. Compared with the control and hAMSC-exos, E-exos treatment downregulated the expression of Notch-1, Notch-4, and Jagged-1 (Figure 5). These data suggest that Notch signaling is affected by E-exos, potentially by the efficient delivery of miR-187-3p by the electroporated exosomes, as indicated by Notch signaling activation not being higher in the E-exos (NC)-treated group than it was in the group treated with naïve exosomes.
Discussion
Exosomes are known as natural nanovesicles and have roles in intercellular communication and molecular transfer. They have been studied for several years (24,25). The ability of exosomes to transfer bioactivators, such as cytokines, signaling lipids, and miRNAs, between cells makes them promising carriers for the effective therapeutic delivery of compounds for the treatment of various conditions, including cancer (26), inflammation (27), and infection (28).
MSC-derived exosomes can be engineered to shuttle miRNAs to target cells. For example, exosomes from miR-199a-expressing MSCs have been shown to be able to deliver miR-199a to glioma cells, resulting in the suppression of glioma cell proliferation, invasion, and migration (29). In addition, miR-644-5p carried by MSC-derived exosomes was found to target p53 regulation to inhibit ovarian granulosa cell apoptosis (30). Also, in a rat model of calvarial defect, exosomes derived from miR-375-overexpressing hAMSCs enhanced bone regenerative capacity (31). In the present study, we observed that miR-187-3p-loaded hAMSC-exos decreased HemSC viability by inactivating Notch signaling via their targeting of Notch-1/Notch-4/Jagged-1 in vitro. Also, free miR-187-3p or naïve hAMSC-exos partially inhibited the viability of HemSCs, but E-exos significantly inhibited cell viability. These results indicate that the exosome-based delivery strategy was crucial for miR-187-3p to function more effectively in the intracellular environment. Moreover, due to its natural lipid bilayer membrane, exosomes protected the miRNA against degradation by the physical environment and increased the intracellular uptake by target cells (32).
miRNAs are small, non-coding regulatory RNA molecules that function as post-transcriptional gene regulators of various cell behaviors, including proliferation, apoptosis, and angiogenesis. Changes in miRNA expression patterns can contribute to many physiological and pathological events, including several cancers (33-35). However, in the extracellular environment, free miRNAs are susceptible to hydrolyzation by RNase. To protect miRNA against degradation during delivery in the present study, hAMSC-exos were engineered to deliver miR-187-3p. We selected hAMSC-exos as the carrier of exogenous miR-187-3p because hAMSCs have the potential to produce a large number of exosomes. We found that miR-187-3p-loaded E-exos could be taken up by the target cells. Subsequently, E-exos mediated target gene repression. These results are consistent with previous findings that MSC-derived exosomes can be used as nanocarriers to deliver miRNA (36,37). In this context, encapsulating active agents into hAMSC-exos can further enhance their treatment effects, which enhances the promise of hAMSC-exos-based therapy.
Notch signaling is an evolutionarily highly conserved signaling pathway that regulates cell fate decisions in pathological angiogenesis (38). Aberrant activity of the Notch pathway is related to the initiation and progression of some malignancies, although Notch can be oncogenic or tumor suppressive depending on the cellular context. Therefore, Notch signaling represents a significant target for therapeutic agents precisely designed to treat many types of tumors. In contrast with Cui and Shi’s osteosarcoma study, which reported that miR-187 inhibited tumor growth and invasion via direct targeting of mitogen-activated protein kinase (MAPK) 12 (39), the present study found that E-exos loading with miR-187-3p inhibited HemSC viability via inactivation of the Notch pathway (Figure 5). Further analysis revealed that E-exos inactivated Notch signaling by downregulating the expression of Notch-1/Notch-4/Jagged-1 in HemSCs in vitro. Notch proteins are dynamically expressed and Notch signaling is activated in IH cells (21,40). We found that the inhibitory effect of E-exos on HemSC viability could be rescued by a Notch signal inhibitor, suggesting that the Notch-1/Notch-4/Jagged-1 signal plays a critical role in the viability of HemSCs. Furthermore, this study found that naïve hAMSC-exos also negatively affected HemSCs viability, which may be attributed to the diversity of cellular signaling effectors encapsulated in the naïve hAMSC-exos cargo. In the present study, miR-187-3p was integrated into the cargo of naïve hAMSC-exos, which not only downregulated the expression of Notch pathway-related proteins in target cells, but also regulated tumor growth and angiogenesis via other possible pathways (10,41).
In conclusion, the results of the present study demonstrate that miR-187-3p carried by hAMSC-derived exosomes can inhibit HemSC viability by targeting Notch-1/Notch-4/Jagged-1 in the cells. Therefore, miR-187-3p may have potential in the treatment of IH.
Acknowledgments
Funding: This study was financially supported by the grants of the National Natural Science Foundation of China (Nos. 81771087, 81901021 and 81801023).
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://atm.amegroups.com/article/view/10.21037/atm-21-4138/rc
Data Sharing Statement: Available at https://atm.amegroups.com/article/view/10.21037/atm-21-4138/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://atm.amegroups.com/article/view/10.21037/atm-21-4138/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Institute Review Board of Shanghai Ninth People’s Hospital (No. SH9H-2019-T272-1) and informed consent was taken from the parents or guardians of all the patients.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Léauté-Labrèze C, Harper JI, Hoeger PH. Infantile haemangioma. Lancet 2017;390:85-94. [Crossref] [PubMed]
- Darrow DH, Greene AK, Mancini AJ, et al. Diagnosis and Management of Infantile Hemangioma. Pediatrics 2015;136:e1060-104. [Crossref] [PubMed]
- Barros FM, Carneiro F, Machado JC, et al. Exosomes and Immune Response in Cancer: Friends or Foes? Front Immunol 2018;9:730. [Crossref] [PubMed]
- Yeo RW, Lai RC, Zhang B, et al. Mesenchymal stem cell: an efficient mass producer of exosomes for drug delivery. Adv Drug Deliv Rev 2013;65:336-41. [Crossref] [PubMed]
- Engels BM, Hutvagner G. Principles and effects of microRNA-mediated post-transcriptional gene regulation. Oncogene 2006;25:6163-9. [Crossref] [PubMed]
- Vignard V, Labbé M, Marec N, et al. MicroRNAs in Tumor Exosomes Drive Immune Escape in Melanoma. Cancer Immunol Res 2020;8:255-67. [Crossref] [PubMed]
- Strub GM, Kirsh AL, Whipple ME, et al. Endothelial and circulating C19MC microRNAs are biomarkers of infantile hemangioma. JCI Insight 2016;1:e88856. [Crossref] [PubMed]
- Mong EF, Akat KM, Canfield J, et al. Modulation of LIN28B/Let-7 Signaling by Propranolol Contributes to Infantile Hemangioma Involution. Arterioscler Thromb Vasc Biol 2018;38:1321-32. [Crossref] [PubMed]
- Zeng Z, Liu S, Cai J, et al. miR-501 promotes hemangioma progression by targeting HOXD10. Am J Transl Res 2019;11:2439-46. [PubMed]
- Dou C, Liu Z, Xu M, et al. miR-187-3p inhibits the metastasis and epithelial-mesenchymal transition of hepatocellular carcinoma by targeting S100A4. Cancer Lett 2016;381:380-90. [Crossref] [PubMed]
- Casanova-Salas I, Masiá E, Armiñán A, et al. MiR-187 Targets the Androgen-Regulated Gene ALDH1A3 in Prostate Cancer. PLoS One 2015;10:e0125576. [Crossref] [PubMed]
- Ozdemir D, Feinberg MW. MicroRNAs in diabetic wound healing: Pathophysiology and therapeutic opportunities. Trends Cardiovasc Med 2019;29:131-7. [Crossref] [PubMed]
- Bobrie A, Colombo M, Krumeich S, et al. Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J Extracell Vesicles 2012; [Crossref] [PubMed]
- Zhang L, Mai HM, Zheng J, et al. Propranolol inhibits angiogenesis via down-regulating the expression of vascular endothelial growth factor in hemangioma derived stem cell. Int J Clin Exp Pathol 2014;7:48-55. [PubMed]
- Wu X, Showiheen SAA, Sun AR, et al. Exosomes Extraction and Identification. Methods Mol Biol 2019;2054:81-91. [Crossref] [PubMed]
- Oosthuyzen W, Sime NE, Ivy JR, et al. Quantification of human urinary exosomes by nanoparticle tracking analysis. J Physiol 2013;591:5833-42. [Crossref] [PubMed]
- Lässer C, Eldh M, Lötvall J. Isolation and characterization of RNA-containing exosomes. J Vis Exp 2012;e3037. [PubMed]
- Alvarez-Erviti L, Seow Y, Yin H, et al. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol 2011;29:341-5. [Crossref] [PubMed]
- Coumans FAW, Brisson AR, Buzas EI, et al. Methodological Guidelines to Study Extracellular Vesicles. Circ Res 2017;120:1632-48. [Crossref] [PubMed]
- Koritzinsky EH, Street JM, Star RA, et al. Quantification of Exosomes. J Cell Physiol 2017;232:1587-90. [Crossref] [PubMed]
- Wu JK, Adepoju O, De Silva D, et al. A switch in Notch gene expression parallels stem cell to endothelial transition in infantile hemangioma. Angiogenesis 2010;13:15-23. [Crossref] [PubMed]
- Wu JK, Kitajewski JK. A potential role for notch signaling in the pathogenesis and regulation of hemangiomas. J Craniofac Surg 2009;20:698-702. [Crossref] [PubMed]
- Edwards AK, Glithero K, Grzesik P, et al. NOTCH3 regulates stem-to-mural cell differentiation in infantile hemangioma. JCI Insight 2017;2:93764. [Crossref] [PubMed]
- Valadi H, Ekström K, Bossios A, et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 2007;9:654-9. [Crossref] [PubMed]
- Liang X, Zhang L, Wang S, et al. Exosomes secreted by mesenchymal stem cells promote endothelial cell angiogenesis by transferring miR-125a. J Cell Sci 2016;129:2182-9. [Crossref] [PubMed]
- Kalluri R. The biology and function of exosomes in cancer. J Clin Invest 2016;126:1208-15. [Crossref] [PubMed]
- McDonald MK, Tian Y, Qureshi RA, et al. Functional significance of macrophage-derived exosomes in inflammation and pain. Pain 2014;155:1527-39. [Crossref] [PubMed]
- Schorey JS, Cheng Y, Singh PP, et al. Exosomes and other extracellular vesicles in host-pathogen interactions. EMBO Rep 2015;16:24-43. [Crossref] [PubMed]
- Yu L, Gui S, Liu Y, et al. Exosomes derived from microRNA-199a-overexpressing mesenchymal stem cells inhibit glioma progression by down-regulating AGAP2. Aging (Albany NY) 2019;11:5300-18. [Crossref] [PubMed]
- Sun B, Ma Y, Wang F, et al. miR-644-5p carried by bone mesenchymal stem cell-derived exosomes targets regulation of p53 to inhibit ovarian granulosa cell apoptosis. Stem Cell Res Ther 2019;10:360. [Crossref] [PubMed]
- Chen S, Tang Y, Liu Y, et al. Exosomes derived from miR-375-overexpressing human adipose mesenchymal stem cells promote bone regeneration. Cell Prolif 2019;52:e12669. [Crossref] [PubMed]
- Qiu Y, Li P, Zhang Z, et al. Insights Into Exosomal Non-Coding RNAs Sorting Mechanism and Clinical Application. Front Oncol 2021;11:664904. [Crossref] [PubMed]
- Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer 2015;15:321-33. [Crossref] [PubMed]
- Li J, Zhang Y, Liu Y, et al. Microvesicle-mediated transfer of microRNA-150 from monocytes to endothelial cells promotes angiogenesis. J Biol Chem 2013;288:23586-96. [Crossref] [PubMed]
- Zhuang G, Wu X, Jiang Z, et al. Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK-STAT pathway. EMBO J 2012;31:3513-23. [Crossref] [PubMed]
- Qiu G, Zheng G, Ge M, et al. Mesenchymal stem cell-derived extracellular vesicles affect disease outcomes via transfer of microRNAs. Stem Cell Res Ther 2018;9:320. [Crossref] [PubMed]
- Pakravan K, Babashah S, Sadeghizadeh M, et al. MicroRNA-100 shuttled by mesenchymal stem cell-derived exosomes suppresses in vitro angiogenesis through modulating the mTOR/HIF-1α/VEGF signaling axis in breast cancer cells. Cell Oncol (Dordr) 2017;40:457-70. [Crossref] [PubMed]
- Rehman AO, Wang CY. Notch signaling in the regulation of tumor angiogenesis. Trends Cell Biol 2006;16:293-300. [Crossref] [PubMed]
- Cui C, Shi X. miR-187 inhibits tumor growth and invasion by directly targeting MAPK12 in osteosarcoma. Exp Ther Med 2017;14:1045-50. [Crossref] [PubMed]
- Ji Y, Chen S, Xiang B, et al. Jagged1/Notch3 Signaling Modulates Hemangioma-Derived Pericyte Proliferation and Maturation. Cell Physiol Biochem 2016;40:895-907. [Crossref] [PubMed]
- Dong F, Zhang Y, Xia F, et al. Genome-wide miRNA profiling of villus and decidua of recurrent spontaneous abortion patients. Reproduction 2014;148:33-41. [Crossref] [PubMed]


