
Clinical characteristics of recovered COVID-19 patients with re-detectable positive RNA test
Introduction
In December 2019, lots of unexplained pneumonia cases appeared which clinical manifestations suggested viral pneumonia. Deep sequencing of lower respiratory tract samples identified a novel coronavirus named the severe acute respiratory syndrome Coronavirus-2 (SARS-CoV-2), and the disease it caused was named coronavirus disease 2019 (COVID-19) (1). Over 300,000 cases of COVID-19 were reported from December 2019 to March 23, 2020. COVID-19, has resulted in 13,000 deaths, globally (1), with The World Health Organization declaring this virus as a pandemic (2). Generally, COVID-19 is less severe and fatal than the SARS. However, some patients, especially the elderly with co-morbidities are prone to developing more severe symptoms and require urgent medical interventions (3,4). Many literature reports have retrospectively analyzed the clinical characteristics of patients infected with SARS-CoV-2Jeny (3-8). Recently, an increasing number of patients with COVID-19 were discharged from the hospital and received regular follow-up and observation. Re-detectable positive (RP) diagnoses of SARS-CoV-2 RNA test in some recovered patients have been reported (9-13). The first reported four patients with COVID-19 who met criteria for hospital discharge or discontinuation of quarantine in China were reported with positive SARS-CoV-2 RNA test 5 to 13 days later which suggest that at least a proportion of recovered patients still may be virus carriers (9). Parts of pediatric patients remained positive viral RNA in stools for longer than 4 weeks which indicates the potential for the virus to be transmitted through fecal excretion and detection of fecal SARS-CoV-2 RNA would be more sensitive (12). Three readmission patients of COVID-19 with negative IgM and positive IgG was described with no fever symptoms and absorbed lesions on computed tomography (CT) images which suggested serum antibody responsive to SARS-CoV-2 associated the viral shedding (13). The management of RP patients has attracted wide attention. However, the RP population reported in the literature was small, with a short duration of follow-up. In addition, the clinical characteristics, potential impact, and significance of RP patients remain unknown, making it difficult to provide empirical information and evidence-based support for the management of patients with COVID-19 during recovery.
This study retrospectively analyzed the clinical characteristics of 38 RP patients and 224 non-RP (NRP) patients who recovered from COVID-19. It was revealed that RP patients were characterized by younger age and milder conditions. They also displayed minor symptoms, more sustained remission by CT imaging, earlier RNA negative conversion, and similar plasma antibody levels during their hospitalization period. They showed no obvious disease progression and infectivity when re-admitted to the hospital. The hyper-sensitive detection method which identified SARS-CoV-2 RNA molecules from most samples initially confirmed as RNA-negative by a commercial kit suggests that recovering COVID-19 patients were potential carriers during this period. These findings provide key information for the effective management of COVID-19 patients during their convalescent phase.
We present the following article in accordance with the MDAR reporting checklist (available at http://dx.doi.org/10.21037/atm-20-5602).
Methods
Study design and participants
A total of 262 confirmed COVID-19 patients discharged from Shenzhen Third People’s Hospital from January 23, 2020 to February 25, 2020 were enrolled in this study. All discharged patients continued to be isolated and were observed for 14 days. Weekly follow-up in person and SARS-CoV-2 RNA detection were also performed at the same time. Patients were subsequently followed-up for at least an additional two weeks, post isolation. Unlike recovered NRP patients, who were closely followed-up outside the hospital, individuals identified as RP were re-admitted to hospital for further medical observation. The close contacts of RP patients also underwent weekly follow-up in person for 14 day. This study was approved by the Ethics Committee of The Third People’s Hospital of Shenzhen (2020-115), which waived the requirement for written patient consent for this retrospective analysis. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). All patients provided oral consent to participate in this retrospective study.
Clinical definition
According to the guideline for the diagnosis and treatment for novel coronavirus pneumonia (the sixth edition), published by National Health Commission of the People’s Republic of China (14), all initial diagnoses of COVID-19 were confirmed with positive respiratory RT-PCR tests. The discharge criteria of the recovered patients included: temperature returned to normal for more than 3 days, respiratory symptoms significantly improved, and significant absorption of pulmonary lesions by chest CT imaging, and a minimum of two consecutive negative RNA test results, performed at least 24 hours apart. RP patients were confirmed by digestive (anal swab) and respiratory positive RT-PCR tests. Since February 22, 2020, evaluation of negative anal swab was supplemented for the discharge criteria in Shenzhen Third People’s Hospital.
Data collection
The medical records of 262 patients, recovered from COVID 19 were reviewed. These records also comprised of 38 RP patients. The epidemiological, demographic, clinical, laboratory data of the patients were collected, summarized and analyzed. According to the initial chest CT imaging post admission, the severity of pulmonary inflammation was divided into mild, moderate and severe, based on lesions involving unilateral lobe, multiple lobes in both lungs, and all lobes in both lungs, respectively. The remission of the lesions was evaluated by chest CT within 7 days after admission. Temporary progression was indicated by increased lesions, whilst persistent remission was indicated by stable, absorbed or decreased lesions.
qRT-PCR and Sherlock assay for SARS-CoV-2 RNA detection
The quantitative reverse transcription polymerase chain reaction (qRT-PCR) was assessed as described previously (15). Nasopharyngeal and anal specimens collected during hospitalization were sent to the laboratory in a viral transport case. Total nucleic acid extraction from the samples was performed using the QIAamp RNA Viral Kit (Qiagen, Heiden, Germany). Quantitative RT-PCR was undertaken using a China Food and Drug Administration (CFDA) approved commercial kit specific for 2019-nCoV detection (GeneoDX Co., Ltd., Shanghai, China) or Sherlock kit, gifted from Feng Zhang lab according to the manual. Each RT-PCR assay provided a Ct value, representing the number of cycles required for the fluorescent signal to cross the threshold for a positive test. A higher Ct value is correlated with a lower viral load. Specimens were considered positive if the Ct value was ≤37.0, and negative if the viral load was undetectable. Specimens with a cycle-threshold value higher than 37 were repeated. The specimen was considered positive if the repeat results were equivalent to the initial result and between 37 and 40. If the repeat Ct was undetectable, the specimen was considered negative. All procedures involving clinical specimens and SARS-CoV-2 were performed in a biosafety level 3 laboratory. Meanwhile, next-generation sequencing of samples were performed from samples of three patients.
CMIA assay for anti-SARS-CoV-2 antibodies
The SARS-CoV-2 specific total antibody (Ab), IgG, IgA, and IgM in plasma was tested using a Chemiluminescence Microparticle Immuno Assay (CMIA). Briefly, recombinant SARS-CoV-2-RBD (Receptor binding domain) proteins were immobilized, and HRP-conjugated antigen detected total antibodies, IgM, IgA and IgG. Total antibodies were tested using double-antigens sandwich enzyme-linked immunosorbent assay (Ab-ELISA). IgM was tested by the IgM µ-chain capture method (IgM-ELISA). IgA and IgG were tested using indirect ELISA. The testing kits were supplied by the Beijing Wantai Biological Pharmacy Enterprise Co., Ltd., China. The relative fluorescence of sample to control (COI) was used to measure the antibody concentration. The larger the COI value, the higher the concentration. When COI exceeded 1, the result was deemed positive.
Statistical analysis
Statistical analysis was performed using SPSS 26.0 (IBM, Chicago). All tests were two-sided, and a P<0.05 was considered significant. Continuous variables were evaluated using the median and interquartile range (IQR) values. Chi-square or Fisher exact tests were utilized to compare the proportions of the categorical variables.
Results
Demographic, epidemiological and clinical characteristics
A total of 262 patients were discharged from January 23, 2020 to February 25, 2020 and weekly followed-up in person with detection of SARS-CoV-2 RNA in nasopharyngeal and anal specimens for at least 14 days. Mild, moderate and severe patients accounted for 11.4% (n=30), 81.0% (n=212) and 7.6% (n=20), respectively in this study population. Up to March 10, 14.5% of convalescent patients (n=38) were re-detected as SARS-CoV-2 RNA positive during their follow-up period. Severe patients were found to be RNA negative (Table S1).

Full table
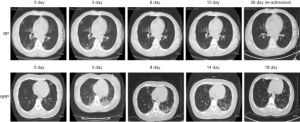
Our findings revealed that the vast majority of RP patients (94.7%, n=36) were younger than 60 years of age. Among them, patients younger than 14 years old were more prevalent compared to those aged between 14 and 60 years (35.0% vs. 16.0%, P<0.01) (Table 1). In addition, the proportion of RP patients displaying mild symptoms (36.7%) was significantly higher than what was observed in NRP patients (12.7%, 19/204, P<0.01, Table S1). No significant difference in gender distribution was exhibited between RP and NRP patients. Notably, the incidence of fever as part of the initial symptoms was higher in mild NRP patients, compared to RP patients (P<0.01). Additionally, 45.5% of mild RP patients displayed only upper respiratory symptoms at the first admission, unlike mild NRP patients, who exhibited lower respiratory symptoms (Table 1). Initial lesion analysis of the chest by CT scan showed no difference in the extent of lesions between moderate RP and NRP patients. However, the incidence of RP (85.2%) was closely related to sustained remission, as indicated by CT imaging of the chest, compared to NRP patients, where 36.2% displayed transient progression during their initial hospitalization (Table 1 and Figure 1). There was no significant difference in the administration of steroid and antiviral therapies between RP and NRP patients during their first hospitalization. In addition, RP patients did not exhibit a higher incidence traveling and living in Hubei province compared to NRP patients.
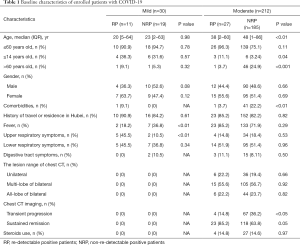
Full table
Differential RNA dynamic in RP and NRP patients
No differences were noted in the duration from illness onset to RNA negative-conversion and hospitalization days between RP and NRP patients. Analysis of the duration between illness onset and RNA negative-conversion was performed. The study revealed that RNA negative conversion occurred mostly within 2–3 weeks from the onset of illness in 63.6% of mild, and 1–2 weeks in 22.2% moderate RP patients. By contrast, there were more NRP patients who displayed RNA negative-conversion after 3 weeks from illness onset, regardless of disease severity (Table 2). The data showed that early RNA-negative conversion was characteristic of RP patients, while NRP patients exhibited late viral clearance.
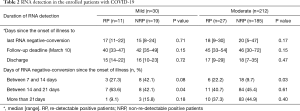
Full table
Changes in serum anti-SARS-CoV-2 antibodies
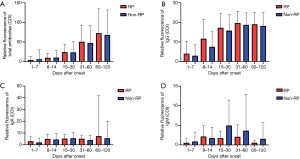
To evaluate the effect of serum-specific antibody levels on the incidence of RP, we analyzed the difference in anti-SARS-CoV-2 antibody levels in RP and NRP patients. A total of 989 plasma samples, including those from 36 RP and 228 NRP patients were examined for total Ab, IgM, IgG, and IgA against SARS-CoV-2. Total antibodies (Figure 2A), IgG (Figure 2B) and IgA (Figure 2C) were detected four months or longer at sustained levels in both groups. No differences in antibody levels were observed between RP and NRP patients. IgM levels increased slowly before exhibiting a decline in both the groups. Peak IgM levels (Figure 2D) in RP patients was slightly higher than NRP patients 15–30 days after illness onset, with a P value (0.056) exceeding 0.05.
Supplementing negative results via anal swab test at discharge failed to reduce RP occurrence of COVID-19 patients
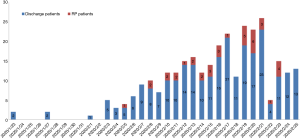
To determine if increasing site tests at discharge affected the incidence of RP, we compared its occurrence before and after the introduction of a supplementary test. The negative anal swab detection was an added procedure as part of the discharge criterion on 22nd February, 2020 in COVID-19 patients. Our results showed that there was no statistical difference in the occurrence of RP patients before or after February 22, 2020 (14.5% vs. 14.3%, P=0.77, Figure 3). The data indicated that supplementary testing at additional detection sites of SARS-CoV-2 RNA failed to reduce the incidence of RP in convalescent patients.
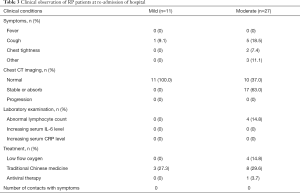
RP patients showed no obvious clinical symptoms and disease progression
All 38 RP patients were re-admitted to hospital for further medical observation. The analysis showed that these patients did not display fever, and a small number of patients reported mild cough and chest tightness, which were previously observed and did not increase in severity (Table 3). All patients recovered from mild conditions (n=30). CT imaging revealed 37.0% of moderate patients displayed normal chests with no signs of inflammation. The remaining 63.0% (n=17) of patients in the same group recovered from moderate conditions and displayed stable or reduced chest inflammation, as confirmed by CT imaging (Figure 1). All RP patients exhibited lymphocyte, plasma IL-6 and CRP levels within the normal range of the upon admission. Only 1 patient received transient interferon-alpha inhalation therapy, and 4 patients received low-flow oxygen inhalation therapy and 11 patients received traditional Chinese medicine after admission (Table 3).
Full table
All the convalescent patients with COVID-19 in our cohort were required to be isolated at home or undergo intensive isolation, yielding only 21 close contacts. Up to March 10, 2020, all close contacts tested negative for SARS-CoV-2 RNA, and no suspicious clinical symptoms were reported in those close contacts (Table 3).
Hyper-sensitive methods potentially improved SARS-CoV-2 RNA detection in RP patients
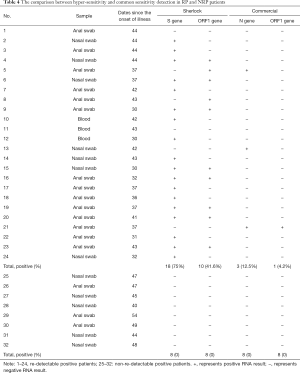
To investigate the possibility of false negative diagnoses due to the low sensitivity of commercial RNA detection kits, an alternative method with higher detection potential was applied to Anal swab, Blood and Nasal swab from RP and NRP patients displaying similar illness days. Twenty-four samples were obtained from 15 RP patients, 5–7 days since the onset of the re-admission. Our findings revealed that 75% of spike genes and 41.6% ORF1 genes were detected using this hyper-sensitive method on, in contrast to 12.5% N genes and 4.2% ORF1 genes, identified by the commercial detection kit. Eight of 15 RP patients were confirmed as RNA positive using the hyper-sensitive kit, whereas only 1 person tested positive with the commercial kit. By contrast, 8 samples from NRP patients were detected to be negative by both methods (Table 4). The data demonstrates that hyper-sensitive methods potentially improved RNA positive detection in RP patients.
Full table
Discussion
Several studies have confirmed the existence of RP patients (9-13), however their clinical characterization was not well defined or with limited clinical samples. This retrospective study analyzed the clinical and follow-up data in a cohort of RP and NRP patients during the same discharge period. Up to March 10, 2020, 38 RP patients were present, accounting for 14.5% of discharged patients during the same follow-up period. These patients displayed several significant features, including younger age and mild and/or moderate symptoms during hospitalization, which is consistent with previous reports (9,12). Mild RP patients were usually below 14 years of age and moderate RP patients were below the age of 60. By contrast, no severe patients were found to be RP within a similar follow-up period. In addition, more RP patients displayed minor symptoms during hospitalization such as fewer comorbidities, less severe fever, and increased upper respiratory symptoms. CT imaging indicated that RP patients had no lesions or maintained sustained remission in their lungs as those of NRP patients. The data indicated that RP patients were characterized by a younger age and minor symptoms during hospitalization.
Viral load is usually considered to be related to disease outcome (15,16). The present study indicated that RNA negative-conversion commonly occurred 2–3 weeks from the onset of illness in moderate RP patients, compared to over 3 weeks in NRP patients of the same disease severity. The significantly shortened RNA negative-conversion duration may affect the persistence of high adaptive immunity levels (17). Our recent study indicated that a higher antibody titer in the plasma was independently associated with disease severity in patients with COVID-19 (18). However, RP and NRP patients displayed similar plasma IgG and IgM levels. Future studies are required to investigate host immune responses and its effects on determining clinical outcomes in viral infection (19,20).
We also comprehensively characterized the clinical symptoms of RP patients when re-admitted to the hospital. No obvious clinical evidence of disease progression or recurrence was found in these patients from CT imaging and laboratory tests. Additionally, no antibiotics, steroids, antiviral agents and continuous supplemental oxygenation were required. RP patients also displayed significantly reduced inflammatory response during re-admission. The data indicated that the diseases of RP patients did not increase in severity despite the positive RNA detection for SARS-CoV-2. More importantly, these patients did not be reinfected after discharge, consistent with a recent longitudinal study in SARS-CoV-2 infected rhesus macaques, where reinfection could not occur in convalescent monkeys (21). Long-term follow-up of close contacts with RP patients will warrant the evaluation of the potential risk of RP.
The mechanisms underlying RP occurrence remain unclear. The possible reasons proposed by a large number of experts are related to several virological, immunological and sampling methodological factors. From a viral standpoint, false negatives (22), viral residual (12), intermittent viral release (12) and viral distribution (23,24) are usually considered as major factors. Our data support the notion that the false negative diagnoses originating from commercial kits may partially account for RPs, as the kits only provide a 30–50% positive rate of detection (24,25). In 24 samples from RP patients, RNA was detected as negative for both N gene and ORF1b gene several days after hospital re-admission using commercial kit, whose lower limit of detection (LOD) was relatively high (500 copies/mL). However, a higher sensitivity Sherlock kit with an LOD of 100 copies/mL (26), confirmed 75% of samples as positive for the S gene and 41.6% for ORF genes. The commercial kit therefore failed to identify half of positive subjects within the RP patient population during hospitalization. By contrast, the Sherlock or commercial kit did not detect any of the 8 samples from NRP patients as positive. However, the Sherlock kit detected a sample previously confirmed by SRAS-CoV-2 sequencing, as positive (data not shown). Therefore, future studies are required with the aim to improve the sensitivity and specificity of detection kits to facilitate the accurate diagnosis of clinical samples. The data indicated that false positive results, detected by the current commercially available kit may account for the occurrence of RP patients, to some extent.
Another viral factor pertains to long-term virus residual levels in the gut and other tissues, similar to SARS (27). A recent study indicated that SARS-CoV-2 nucleic acid can persist in the digestive tract and feces for nearly 50 days (28). Thus, extending the follow-up time is necessary for the COVID-19 patients when they were discharged. However, our results indicated that additional discharge criterion such as negative results from supplementary RNA testing, which originated from anal swabs, did not significantly reduce the incidence of RP patients. Thus other factors may be associated with the RP patients. We could not exclude sampling methodological factors including differential sampling and operational methods, sample quality, and technician expertise levels. Nor could we exclude immunological factors including low mucosal immune responses such as low IgA levels. These factors may take some uncertain risks leading the occurrence of RP patients (4,28). Future studies focused on the application of the hyper-sensitive detection kit, combined with the detection of multiple samples and more immune markers in should reduce the incidence of RP.
This study has several limitations. Firstly, this study was a single-center retrospective study with a short follow-up duration. More clinical observations are needed to evaluate the potential risk of SARS-CoV-2 recurrence and infection. Secondly, the dynamics of SARS-CoV-2 RNA in COVID-19 patients need to be monitored and evaluated for RP. Furthermore, additional studies which measure the dynamic changes of serum specific antibody levels in RP patients and evaluate the continuous protective effect of serum specific antibodies on patients with COVID-19 are required. Finally, distinctions are required between RP, relapsed, and convalescent patients, where two distinct prevention and control strategies will be adopted.
Collectively, our findings revealed the clinical features of RP patients who did not exhibit the recurrence of clinical symptoms and abnormal laboratory tests. However, hyper-sensitive detection methods revealed the existence of SARS-CoV-2 RNA in RP patient specimens, initially tested as negative using the commercial kit. Therefore, the development of a more accurate kit is required for quantitative assessments of the RNA dynamics and additional discharge criteria to aid in the guidance of clinical decisions associated with COVID-19. This study provided valuable empirical information and clinical evidence support for effective management of COVID-19 patients during the convalescent period. Further study should evaluate the potential clinical significance and transmission risk of RP patients.
Acknowledgments
We acknowledge the work and contribution of all the health providers from Shenzhen Third People’s Hospital.
Funding: This research was funded by the Medical and Health Three Famous Projects in Shenzhen and awarded to Professor Liu You-Ning, General Hospital of the Chinese People’s Liberation Army, Respiratory System Critical Illness, and Major Emerging Infectious Diseases Diagnosis and Research Team (SZSM201612025). This research was funded by the 2020 Guangdong Province Special Project on Emergency Research on Prevention and Control of New Coronavirus Infection Technology (2020B1111340030).
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/atm-20-5602
Data Sharing Statement: Available at http://dx.doi.org/10.21037/atm-20-5602
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/atm-20-5602). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was approved by the Ethics Committee of The Third People’s Hospital of Shenzhen (2020-115), which waived the requirement for written patient consent for this retrospective analysis. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). All patients provided oral consent to participate in this retrospective study.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Available online: Health Organization.https://www.who.int/emergencies/diseases/novel-coronavirus-2019.World
- WHO characterizes COVID-19 as a pandemic. Available online: (accessed Mar 14, 2020).https://www.who.int/emergencies/diseases/novel-coronavirus-2019/events-as-they-happen
- Guan WJ, Ni ZY, Hu Y, et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N Engl J Med 2020;382:1708-20. [Crossref] [PubMed]
- Zhou F, Yu T, Du R, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 2020;395:1054-62. [Crossref] [PubMed]
- Wang M, Wu Q, Xu WZ, et al. Clinical diagnosis of 8274 samples with 2019-novel coronavirus in Wuhan. medRxiv 2020. DOI: . [Crossref]
- Chen N, Zhou M, Dong X, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 2020;395:507-13. [Crossref] [PubMed]
- Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020;395:497-506. [Crossref] [PubMed]
- Wang D, Hu B, Hu C, et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020;323:1061-9. [Crossref] [PubMed]
- Lan L, Xu D, Ye G, et al. Positive RT-PCR Test Results in Patients Recovered From COVID-19. JAMA 2020;323:1502-3. [Crossref] [PubMed]
- Ling Y, Xu SB, Lin YX, et al. Persistence and clearance of viral RNA in 2019 novel coronavirus disease rehabilitation patients. Chin Med J (Engl) 2020;133:1039-43. [Crossref] [PubMed]
- Qu YM, Cong HY. Positive result of SARS-Co-2 in sputum from a cured patient with COVID-19. Travel Med Infect Dis 2020;34:101619. [Crossref] [PubMed]
- Xing Y, Ni W, Wu Q, et al. Prolonged presence of SARS-CoV-2 in feces of pediatric patients during the convalescent phase. medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Wei Fu, Qian Chen, and Tao Wang. Letter to the Editor: Three cases of re-detectable positive SARS-CoV-2 RNA in recovered COVID-19 patients with antibodies. J Med Virol 2020. [Epub ahead of print].
- Available online: (accessed Mar 9, 2020).http://www.nhc.gov.cn/yzygj/s7653p/202003/46c9294a7dfe4cef80dc7f5912eb1989.shtml
- Liu Y, Yang Y, Zhang C, et al. Clinical and biochemical indexes from 2019-nCoV infected patients linked to viral loads and lung injury. Sci China Life Sci 2020;63:364-74. [Crossref] [PubMed]
- Wei L, Ming SQ, Zou B, et al. Viral Invasion and Type I Interferon Response Characterize the Immunophenotypes during Covid-19 Infection. Available online: https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3555695
- Dong C, Ni L, Ye F, et al. Characterization of anti-viral immunity in recovered individuals infected by SARS-CoV-2. MedRxiv 2020. Available online: https://doi.org/ [Crossref]
- Zhao J Jr, Yuan Q, Wang H, et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Guo X, Guo Z, Duan C, et al. Long-Term Persistence of IgG Antibodies in SARS-CoV Infected Healthcare Workers. medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Liu W, Fontanet A, Zhang PH, et al. Two-year prospective study of the humoral immune response of patients with severe acute respiratory syndrome. J Infect Dis 2006;193:792-95. [Crossref] [PubMed]
- Bao L, Deng W, Gao H, et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaque. medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Li D, Wang D, Dong J, et al. False-Negative Results of Real-Time Reverse-Transcriptase Polymerase Chain Reaction for Severe Acute Respiratory Syndrome Coronavirus. Korean J Radiol 2020;21:505-8. [Crossref] [PubMed]
- Xia J, Tong J, Liu M, et al. Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection SARS-CoV-2 through conjunctiva. J Med Virol 2020;92:589-94. [Crossref] [PubMed]
- Wölfel R, Corman VM, Guggemos W, et al. Virological assessment of hospitalized cases of coronavirus disease 2019. medRxiv 2020. https://doi.org/ [Crossref]
- Lin C, Xiang J, Yan M, et al. Comparison of throat swabs and sputum specimens for viral nucleic acid detection in 52 cases of novel coronavirus (SARS-Cov-2) infected pneumonia (COVID-19). medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Zhang F, Abudayyeh OO, Gootenberg JS. A protocol for detection of COVID-19 using CRISPR diagnostics (v.20200321). Available online: https://www.broadinstitute.org/files/publications/special/COVID-19%20detection%20(updated).pdf
- Xiao F, Tang M, Zheng X, et al. Evidence for gastrointestinal infection of SARS-CoV-2. medRxiv 2020. Available online: https://doi.org/ [Crossref]
- Wu Y, Guo C, Tang L, et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol Hepatol 2020;5:434-5. [Crossref] [PubMed]
(English Language Editor: E. Tan)


