
A narrative review of tumor heterogeneity and challenges to tumor drug therapy
Introduction
With the accumulation of mutations in key oncogenes and cancer suppressor genes as well as deletions and translocations, normal cells can undergo enhanced cell proliferation, escape growth inhibition and cell death signals, induce changes such as angiogenesis, and ultimately activate programs that lead to the invasion and metastasis of tissues, and thus to the formation of cancer (1,2). It is not static after its formation, but a dynamic evolutionary process. As the disease evolves, cancers usually show a more pronounced heterogeneity, resulting in tumors that may contain different collections of cells. This heterogeneity will lead to uneven distribution of genetic diversity of tumor cell subpopulations within the lesions and tumors (spatial heterogeneity) and temporal variations in the molecular composition of cancer cells (temporal heterogeneity; Figure 1). The spatial and temporal heterogeneity lays the foundation for tumor drug resistance. Designing more targeted drug combinations based on tumor heterogeneity is necessary to maximize drug effectiveness and minimize toxicity (3). Therefore, the accurate evaluation of tumor heterogeneity is of great clinical significance to enhance the therapeutic efficacy and improve the prognosis.
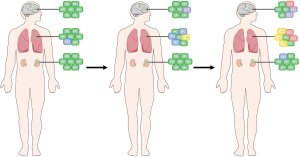
In this review, we summarized the mechanisms and manifestations of tumor heterogeneity and focus on their effects on drug resistance. Finally, we discussed the challenges and solutions brought by tumor heterogeneity to gene detection to improve cancer detection and treatment.
We present the following article in accordance with the Narrative Review reporting checklist (available at https://dx.doi.org/10.21037/atm-21-1948).
Tumor heterogeneity
The earliest evidence that tumors are heterogeneous can trace back to 1833 (Figure 2). Muller and Virchow first put human tumor samples under the microscope and discovered that cancer cells have different morphologies and distinguished cancer subtypes in a single tumor (4). The first evidence that cancer is a genetic disease was the observation of abnormal mitochondria in cancer cells by Hansmann in 1890 (5). Theodor Boveri found in 1905 that abnormal mitoses were closely associated with the development of malignancy (6). In the 20th century, the development of cell biology helped researchers to explore tumors further. Makino et al. (7) found differences in functional and genetic information between spontaneous tumors and normal cells, which also confirmed the heterogeneity of tumors. In the latter decades of the 20th century, several hypotheses were proposed to explain the mechanisms of tumor heterogeneity formation. Nowell first proposed the theory of clonal evolution in 1976 (8). Harris et al. (9) proposed the concept of metastatic variants. With the continuous development and gradual improvement of technologies such as next-generation sequencing (10) and single-cell sequencing (11), it has become possible to achieve deep sequencing of individual cells or whole genomes of tumors in different regions spatially or temporally, thus deepening researchers’ understanding of the corresponding theoretical and molecular mechanisms of tumor heterogeneity.
The mechanisms of tumor heterogeneity
Tumor heterogeneity is reflected not only in time and space but also in structural heterogeneity, gene heterogeneity, protein heterogeneity, functional heterogeneity, and so on. Generally speaking, the main factors affecting tumor heterogeneity include internal cell factors and cell microenvironment. The specific mechanisms can be categorized into genomic instability, epigenetic modifications, plastic gene expression, and different microenvironments (Figure 3). Different mechanisms primarily mediate heterogeneity in various types of tumors and, as tumors develop, mechanisms from multiple sources of heterogeneity may interact and work together, ultimately leading to heterogeneous cancer production.

Genomic instability is considered to be the most familiar internal driving force in the various mechanisms of tumor heterogeneity. DNA repair, telomere maintenance, DNA replication, and chromosome segregation can lead to extensive and random alterations in any part of the genome (12). There is a much higher rate of genetic mutations in cancer cells compared to normal cells. Among the 12 major categories of cancer, the mutation rate per trillion bases ranged from 0.28 to 8.15 (13). Even if the mutation rate is low, the number of base pairs in the human diploid genome is large (about 6 billion), and random mutations are critical in the development of tumorigenesis.
In addition to gene mutations, epigenetic modifications also contribute to tumor heterogeneity. Changes in epigenetic modifications can remain stable in genetic information and can be transmitted to offspring, while there is no change in the DNA sequence (14). Some studies have confirmed that there is a kind of stem cell in the tumor. Only cancer stem cells (CSCs) are capable of infinite self-renewal and powerful differentiation to initiate and maintain tumor growth (15). Thus, tumor development resembles the tissue differentiation hierarchy of normal stem cells. CSCs generate cellular heterogeneity by epigenetic changes, forming a variety of phenotypic non-tumorigenic cells, which constitute most of the cells in the tumor.
Differential gene expression is also one of the essential mechanisms of tumor heterogeneity. The cell surface antigens of some cancer cells are affected by many factors such as drug therapy, which confirms the mechanism of temporary changes in gene expression. Singer et al. (16) performed an analysis of gene expression in a single embryonic stem cell with the aid of single-molecule RNA-FISH and quantitative time-lapse movies, which proved that random gene expression is a fundamental property of cells in response to changes in their environment, which is common in nature.
Different microenvironments of tumor cells will also lead to heterogeneous expression. The microenvironment that causes this inequality is the result of the joint action of many factors. One of the more important is blood supply (17). The difference in the distance from a single cancer cell to the vascular system leads to different nutritional supply and metabolism of cancer cells, which affects the heterogeneity of tumor cells. Besides, other stromal cells, such as fibroblasts, inflammatory cells, and pluripotent mesenchymal cells, also promote the diversity of cancer cell genotypes and phenotypes through secretions, including cytokines, growth factors, and extracellular matrix (ECM) components. These secretions of stromal cells contribute to their chemotaxis and further influence the tumor microenvironment (18).
The clonal evolution model reasonably explains how internal cellular factors and tumor microenvironment lead to intra-tumor heterogeneity. The model holds that in cancer development, random genetic changes create cell pools, a pool of cells that vary in genetic alterations and growth potential, but only cancer cells suitable for the surrounding environment will survive (19). Other sections are gradually eliminated because they do not adapt to the environment. The existence of this different subclone has been confirmed in many kinds of cancer, including human non-small cell lung cancer (NSCLC). The spatial and temporal maps of tumor cells also support subclonal branching evolution. But it has been suggested that drug resistance status can be reversed. Adaptive therapy can maintain a competitive balance between drug-sensitive and drug-resistant clones, stabilizing tumor burden and response to therapy (20,21). Besides, some targeted drugs can reverse tumor chemotherapy resistance status, improve treatment efficacy, and prolong patient survival (22).
Spatial heterogeneity of tumor
Spatial heterogeneity means that within the primary tumor or between the primary tumor and the metastases, there will be differences in characteristics such as genetic information and cell morphology. In a prospective cohort study, 327 tumor regions from 100 cases of early-stage NSCLC were sequenced and analyzed. It turns out that more than 75% of tumor drivers have changed later in the evolution. Both copy number alterations and mutations in somatic cells were found to be broadly heterogeneous (23). Gerlinger et al. (24) found that only 34% of the mutations identified were consistent in all samples when several sites and metastases were detected in patients with the same primary renal tumor. Therefore, it is difficult to make a comprehensive evaluation of the cancer as a whole through local tissue. The information we get from small samples is likely to be a glimpse of the cancer and does not reflect the real situation of the entire primary and metastatic lesions. In order to overcome the characteristics of spatial heterogeneity, we can consider trying to obtain data from different angles in different locations, and comprehensively interpret the test results. For example, some researchers suggested that at least three different regions of the same tumor should be selected during sampling of surgical resection samples from patients with clear cell renal carcinoma to ensure the accuracy of the five key mutation tests (25). However, most cancers are diagnosed at an advanced stage with little benefit from surgical treatment, and taking multiple biopsy samples by puncture may put the patient’s life at greater risk. Therefore, this multi-regional sampling method is an option for the early screening of tumors. Notably, the margin of multi-regional sampling for single tumors must be limited according to the size of the tumor (23).
Temporal heterogeneity of tumors
The temporal heterogeneity of tumors is mainly manifested by the polyclonal properties of tumors that evolve over time, showing a pronounced dynamism. Most of the current research on the temporal heterogeneity of tumors has come from targeted drug therapy. The genomic complexity of tumors usually increases with the continuation of even drug therapy. For example, in a study using epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKIs) to treat patients with EGFR-sensitive mutations in late-stage NSCLC, the T790M mutation in patient plasma was detected at a fixed time (26). The results showed that the T790M mutation positivity rate increased accordingly with a longer treatment duration. Therefore, the diagnostic results from a single sampling may become outdated quickly. Considering this characteristic of temporal tumor heterogeneity, we can select appropriate biomarkers for dynamic monitoring of tumors to adjust treatment regimens promptly.
Tumor heterogeneity and drug resistance
In terms of composition, solid tumors include not only cancer cells, but also intravascular cells, immune cells, mesenchymal cells, and so on (1). Furthermore, cancer cells within the tumor vary greatly in molecular characteristics (27), which is known as intra-tumor heterogeneity (28,29).
These intercellular differences in molecular characteristics can develop at genomic, gene expression, and post-translational modification stages, but most kinds of cancers can recognize these differential molecules. Currently, cancer therapies ignore these cellular differences and treat cancer as a homogeneous disease. Most of the targeted drugs targeting the abnormal molecular characteristics of cancer cells are based on a single biopsy, which cannot reflect the overall condition of the tumor (30,31). Similar to the immune response, intrinsic drug resistance and adaptive drug resistance are the mainstream explanations for the mechanism of drug resistance. The inherent drug resistance refers to the resistance of the tumor itself to drugs. Tumor heterogeneity leads to the emergence of subclones with different drug sensitivities, and some subclonal populations possess more excellent drug resistance than the initial tumor cells (32). Cytotoxic drugs can destroy most of the non-resistant subclones in heterogeneous tumors, but a small number of resistant subclones dominate and continue to grow and re-form tumors. And adaptive resistance is a drug resistance mechanism developed by tumor cells that are not initially resistant may also evolve a diverse set of resistance mechanisms after targeted therapy. Several studies on the use of EGFR-TKIs in NSCLC leading to the T790M mutation in the EGFR gene demonstrate it (33-35). In addition, the problem of resistance to tumor immunotherapy caused by heterogeneity cannot be ignored. Chimeric antigen receptor (CAR) T-cell therapy for multiple myeloma resulted in the absence of target antigens, which induced immune escape (36). Programmed Cell Death Protein 1 (PD-1) blockade promoted the proliferation of T-reg cells in gastric cancer, leading to suppression of anti-tumor immunity (37). Combining other therapies such as radiotherapy can increase the sensitivity of tumor immunotherapy (38). The heterogeneity of drug resistance related gene mutations further complicated targeted therapy. Drug resistance gene mutations can occur in different patients, such as multiple mutations of G1202R, G1269A, L1196M, I1171T, and F1174C in anaplastic lymphoma kinase (ALK) (39). Co-mutations of multiple drug resistant genes, such as ALK/KRAS co-alterations, can also occur in the same patient (40). Having a profound understanding of the specific drivers of drug resistance in various types of tumors will help us to understand the essence of cancer better and to develop more targeted and effective cancer therapies (41).
Tumor heterogeneity and multidimensional diagnosis
People’s deepening understanding of tumor heterogeneity has benefited from the rapid development and continuous detection technology progress. In order to overcome tumor heterogeneity, the tests performed on tumors also have the characteristics of sample heterogeneity and platform heterogeneity. At present, circulating tumor cells (CTCs) or circulating tumor nucleic acids (ctDNA and ctRNA), used as a source of the bioanalytics for peripheral blood samples, can track treatment status and monitor the emergence of drug resistance. In addition to blood, common types of samples include pleural fluid, ascites, cerebrospinal fluid, and urine. For some patients with unusual tumors, they should be more flexible in choosing the samples to be sent to ensure the test results’ authenticity. There are various detection platforms that can be applied to tumors, such as real-time polymerase chain reaction (qPCR), digital PCR, next generation sequencing (NGS), and single-cell sequencing.
ctDNA was first developed and characterized by its ease of isolation and long-term preservation, which facilitated targeted deep sequencing (42). And circulating free DNA (cfDNA) levels are higher in cancer patients than in normal healthy people, and only a small fraction represents ctDNA (43). The half-life of cfDNA is about 2 hours, which can realize the real-time monitoring of allele frequency changes and track the changes of clones and subclones. It is the best solution to replace the multi-point tissue sampling, break the time and space heterogeneity, and perform comprehensive tumor assessment (44). There are several sequencing platforms that can be used for the analysis of ctDNA, all of which possess some strengths and weaknesses (45). Both qPCR and dPCR are highly sensitive to the genomic changes detected and can be easily applied to clinical workflows with an easy bioinformatic burden. And dPCR is an absolute quantification technology for DNA molecules that enables absolute quantification of starting samples (46). Although the information obtained by a single dPCR test is limited, the detection sensitivity is the highest among various molecular detection platforms. The new NGS platform, optimized for ctDNA analysis, is highly sensitive and efficient in monitoring low-level diseases. A large number of studies have confirmed the clinical value of NGS in tuberculosis (47) cancer (48), and so on. NGS enables massively parallel sequencing, capable of sequencing millions of DNA fragments simultaneously (49). Under the premise of ensuring the sequencing depth, it can grasp more comprehensive tumor driver gene mutations at one time, which is more suitable for newly diagnosed cancer patients. Besides, NGS-based methods are not only highly sensitive for screening for identified mutations but can also cover the entire target genome to detect unidentified mutations. However, NGS-based methods are expensive and require more sophisticated bioinformatics support (Table 1). To design more intelligent bioinformatics software and introduce automation for testing can minimize the disadvantages of NGS-based methods and make them better suited for clinical applications (50). The other bioanalytical source is CTCs. Although CTC and ctDNA platforms are comparable in terms of clinical potential, there are significant differences in the clinical application of these two platforms. Because ctDNA is widely available and easy to collect, it can be successfully detected by a simple biopsy platform. However, CTC platforms require specialized instrumentation to obtain the target cells for detection, and current methods for detecting CTCs lack standardization and have varying detection thresholds. In terms of biological source, CTCs are intact tumor cells, which can provide more heterogeneous information within the lesion and visualize its heterogeneity level (51). Besides, CTCs can be analyzed by single-cell sequencing to detect DNA, RNA, or protein levels, bringing a complete understanding of the evolution of tumor cell cloning (52). Therefore, in response to different clinical needs, selecting an adapted platform can better reduce the testing cost and shorten the diagnosis time on the premise that the test results are authentic and credible.

Full table
Conclusions and future directions
Tumor heterogeneity is affected by many factors consist of internal cell factors and cell microenvironment. Heterogeneity is the main obstacle to effective and individualized treatment of tumors. How to understand the fickle of tumors in multiple dimensions and how to flexibly use a variety of detection methods to capture the changes of tumors at a time when targeted therapy and immunotherapy are rapidly developing can help to improve the design of diagnosis and treatment plans for cancer and benefit millions of patients.
Acknowledgments
The authors wish to acknowledge Wenhui Wu, for her help in checking the English language use of this review.
Funding: This work was supported by a grant of young talents in Shanghai, National Natural Science Foundation of China (81802255), Young Talents in Shanghai (2019 QNBJ), ‘Dream Tutor’ Outstanding Young Talents Program (fkyq1901), Clinical Research Project of Shanghai Pulmonary Hospital (fk18005), Key Discipline in 2019 (oncology), Project of Shanghai Municipal Science and Technology Commission (Project of Municipal Science and Technology Commission), Scientific research project of Shanghai Pulmonary Hospital (fkcx1903), Shanghai Municipal Commission of Health and Family Planning (2017YQ050), Innovation Training Project of SITP of Tongji University, and key projects of leading talent (19411950300). Youth project of hospital management research fund of Shanghai Hospital Association (Q1902037).
Footnote
Reporting Checklist: The authors have completed the Narrative Review reporting checklist. Available at https://dx.doi.org/10.21037/atm-21-1948
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://dx.doi.org/10.21037/atm-21-1948). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell 2011;144:646-74. [Crossref] [PubMed]
- Kleppe M, Levine RL. Tumor heterogeneity confounds and illuminates: assessing the implications. Nat Med 2014;20:342-4. [Crossref] [PubMed]
- Zhao B, Hemann MT, Lauffenburger DA. Intratumor heterogeneity alters most effective drugs in designed combinations. Proc Natl Acad Sci U S A 2014;111:10773-8. [Crossref] [PubMed]
- Balkwill F, Mantovani A. Inflammation and cancer: back to Virchow? Lancet 2001;357:539-45. [Crossref] [PubMed]
- Hansemann D. Ueber asymmetrische Zelltheilung in Epithelkrebsen und deren biologische Bedeutung. Archiv f pathol Anat 1890;119:299-326. Available online:
10.1007/BF01882039 10.1007/BF01882039 - Weinberg RA. In Retrospect: The chromosome trail. Nature 2008;453:725. Available online:
10.1038/453725a 10.1038/453725a - Makino S. Further evidence favoring the concept of the stem cell in ascites tumors of rats. Ann N Y Acad Sci 1956;63:818-30. [Crossref] [PubMed]
- Nowell PC. The clonal evolution of tumor cell populations. Science 1976;194:23-8. [Crossref] [PubMed]
- Harris JF, Chambers AF, Hill RP, et al. Metastatic variants are generated spontaneously at a high rate in mouse KHT tumor. Proc Natl Acad Sci U S A 1982;79:5547-51. [Crossref] [PubMed]
- Song P, Chen SX, Yan YH, et al. Selective multiplexed enrichment for the detection and quantitation of low-fraction DNA variants via low-depth sequencing. Nat Biomed Eng 2021;5:690-701. [Crossref] [PubMed]
- Rodriguez-Meira A, Buck G, Clark SA, et al. Unravelling Intratumoral Heterogeneity through High-Sensitivity Single-Cell Mutational Analysis and Parallel RNA Sequencing. Mol Cell 2019;73:1292-1305.e8. [Crossref] [PubMed]
- Burrell RA, McGranahan N, Bartek J, et al. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 2013;501:338-45. [Crossref] [PubMed]
- Kandoth C, McLellan MD, Vandin F, et al. Mutational landscape and significance across 12 major cancer types. Nature 2013;502:333-9. [Crossref] [PubMed]
- Dupont C, Armant DR, Brenner CA. Epigenetics: definition, mechanisms and clinical perspective. Semin Reprod Med 2009;27:351-7. [Crossref] [PubMed]
- Sreepadmanabh M, Toley BJ. Investigations into the cancer stem cell niche using in-vitro 3-D tumor models and microfluidics. Biotechnol Adv 2018;36:1094-110. [Crossref] [PubMed]
- Singer ZS, Yong J, Tischler J, et al. Dynamic heterogeneity and DNA methylation in embryonic stem cells. Mol Cell 2014;55:319-31. [Crossref] [PubMed]
- Fukumura D, Duda DG, Munn LL, et al. Tumor microvasculature and microenvironment: novel insights through intravital imaging in pre-clinical models. Microcirculation 2010;17:206-25. [Crossref] [PubMed]
- Junttila MR, de Sauvage FJ. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 2013;501:346-54. [Crossref] [PubMed]
- Shackleton M, Quintana E, Fearon ER, et al. Heterogeneity in cancer: cancer stem cells versus clonal evolution. Cell 2009;138:822-9. [Crossref] [PubMed]
- Strobl MAR, West J, Viossat Y, et al. Turnover Modulates the Need for a Cost of Resistance in Adaptive Therapy. Cancer Res 2021;81:1135-47. [Crossref] [PubMed]
- West J, You L, Zhang J, et al. Towards Multidrug Adaptive Therapy. Cancer Res 2020;80:1578-89. [Crossref] [PubMed]
- Echeverria GV, Ge Z, Seth S, et al. Resistance to neoadjuvant chemotherapy in triple-negative breast cancer mediated by a reversible drug-tolerant state. Sci Transl Med 2019;11:eaav0936 [Crossref] [PubMed]
- Jamal-Hanjani M, Wilson GA, McGranahan N, et al. Tracking the Evolution of Non-Small-Cell Lung Cancer. N Engl J Med 2017;376:2109-21. [Crossref] [PubMed]
- Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 2012;366:883-92. [Crossref] [PubMed]
- Sankin A, Hakimi AA, Mikkilineni N, et al. The impact of genetic heterogeneity on biomarker development in kidney cancer assessed by multiregional sampling. Cancer Med 2014;3:1485-92. [Crossref] [PubMed]
- Wang W, Song Z, Zhang Y. A Comparison of ddPCR and ARMS for detecting EGFR T790M status in ctDNA from advanced NSCLC patients with acquired EGFR-TKI resistance. Cancer Med 2017;6:154-62. [Crossref] [PubMed]
- Jia K, He Y, Dziadziuszko R, et al. T cell immunoglobulin and mucin-domain containing-3 in non-small cell lung cancer. Transl Lung Cancer Res 2019;8:895-906. [Crossref] [PubMed]
- Gupta PB, Fillmore CM, Jiang G, et al. Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell 2011;146:633-44. [Crossref] [PubMed]
- Sharma A, Merritt E, Hu X, et al. Non-Genetic Intra-Tumor Heterogeneity Is a Major Predictor of Phenotypic Heterogeneity and Ongoing Evolutionary Dynamics in Lung Tumors. Cell Rep 2019;29:2164-2174.e5. [Crossref] [PubMed]
- Gonzalez de Castro D, Clarke PA, Al-Lazikani B, et al. Personalized cancer medicine: molecular diagnostics, predictive biomarkers, and drug resistance. Clin Pharmacol Ther 2013;93:252-9. [Crossref] [PubMed]
- Cohen P, Alessi DR. Kinase drug discovery--what's next in the field? ACS Chem Biol 2013;8:96-104. [Crossref] [PubMed]
- de Bruin EC, McGranahan N, Mitter R, et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 2014;346:251-6. [Crossref] [PubMed]
- Hata AN, Niederst MJ, Archibald HL, et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat Med 2016;22:262-9. [Crossref] [PubMed]
- Ramirez M, Rajaram S, Steininger RJ, et al. Diverse drug-resistance mechanisms can emerge from drug-tolerant cancer persister cells. Nat Commun 2016;7:10690. [Crossref] [PubMed]
- Yang S, Mao S, Li X, et al. Uncommon EGFR mutations associate with lower incidence of T790M mutation after EGFR-TKI treatment in patients with advanced NSCLC. Lung Cancer 2020;139:133-9. [Crossref] [PubMed]
- Da Vià MC, Dietrich O, Truger M, et al. Homozygous BCMA gene deletion in response to anti-BCMA CAR T cells in a patient with multiple myeloma. Nat Med 2021;27:616-9. [Crossref] [PubMed]
- Kamada T, Togashi Y, Tay C, et al. PD-1+ regulatory T cells amplified by PD-1 blockade promote hyperprogression of cancer. Proc Natl Acad Sci U S A 2019;116:9999-10008. [Crossref] [PubMed]
- Oweida A, Hararah MK, Phan A, et al. Resistance to Radiotherapy and PD-L1 Blockade Is Mediated by TIM-3 Upregulation and Regulatory T-Cell Infiltration. Clin Cancer Res 2018;24:5368-80. [Crossref] [PubMed]
- Katayama R. Drug resistance in anaplastic lymphoma kinase-rearranged lung cancer. Cancer Sci 2018;109:572-80. [Crossref] [PubMed]
- Schmid S, Gautschi O, Rothschild S, et al. Clinical Outcome of ALK-Positive Non-Small Cell Lung Cancer (NSCLC) Patients with De Novo EGFR or KRAS Co-Mutations Receiving Tyrosine Kinase Inhibitors (TKIs). J Thorac Oncol 2017;12:681-8. [Crossref] [PubMed]
- Chen P, Kuang P, Wang L, et al. Mechanisms of drugs-resistance in small cell lung cancer: DNA-related, RNA-related, apoptosis-related, drug accumulation and metabolism procedure. Transl Lung Cancer Res 2020;9:768-86. [Crossref] [PubMed]
- Lo YM, Corbetta N, Chamberlain PF, et al. Presence of fetal DNA in maternal plasma and serum. Lancet 1997;350:485-7. [Crossref] [PubMed]
- Bryzgunova OE, Konoshenko MY, Laktionov PP. Concentration of cell-free DNA in different tumor types. Expert Rev Mol Diagn 2021;21:63-75. [Crossref] [PubMed]
- Murugaesu N, Wilson GA, Birkbak NJ, et al. Tracking the genomic evolution of esophageal adenocarcinoma through neoadjuvant chemotherapy. Cancer Discov 2015;5:821-31. [Crossref] [PubMed]
- Dong L, Meng Y, Sui Z, et al. Comparison of four digital PCR platforms for accurate quantification of DNA copy number of a certified plasmid DNA reference material. Sci Rep 2015;5:13174. [Crossref] [PubMed]
- Quan PL, Sauzade M, Brouzes E. dPCR: A Technology Review. Sensors (Basel) 2018;18:1271. [Crossref] [PubMed]
- Chen P, Sun W, He Y. Comparison of metagenomic next-generation sequencing technology, culture and GeneXpert MTB/RIF assay in the diagnosis of tuberculosis. J Thorac Dis 2020;12:4014-24. [Crossref] [PubMed]
- Tan AC, Lai GGY, Tan GS, et al. Utility of incorporating next-generation sequencing (NGS) in an Asian non-small cell lung cancer (NSCLC) population: Incremental yield of actionable alterations and cost-effectiveness analysis. Lung Cancer 2020;139:207-15. [Crossref] [PubMed]
- Gu W, Miller S, Chiu CY. Clinical Metagenomic Next-Generation Sequencing for Pathogen Detection. Annu Rev Pathol 2019;14:319-38. [Crossref] [PubMed]
- Chen P, Sun W, He Y. Comparison of the next-generation sequencing (NGS) technology with culture methods in the diagnosis of bacterial and fungal infections. J Thorac Dis 2020;12:4924-9. [Crossref] [PubMed]
- Wu J, Hu S, Zhang L, et al. Tumor circulome in the liquid biopsies for cancer diagnosis and prognosis. Theranostics 2020;10:4544-56. [Crossref] [PubMed]
- Lim SB, Di Lee W, Vasudevan J, et al. Liquid biopsy: one cell at a time. NPJ Precis Oncol 2019;3:23. [Crossref] [PubMed]



