Application of next-generation sequencing technologies in Neurology
Introduction
Numerous neurological diseases have well-established evidence of genetic contributions (1). Recent advances in two main fields of genome technologies: genome-wide association studies (GWAS) and next-generation sequencing (NGS) technology, have led to huge progress in understanding the genetic causes of these diseases. GWAS is based on the common disease-common variant hypothesis, and could provide information on how common genetic variability confers risk for the common diseases (2). While for rare Mendelian disorders, NGS could pinpoint novel genes that contain mutations underlying the phenotype. Most of the neurological diseases, including Alzheimer’s disease (AD), Parkinson’s disease (PD), multiple sclerosis (MS), as well as amyotrophic lateral sclerosis (ALS), are suitable for both approaches: sporadic cases are amenable to GWAS, whereas cases presenting a positive family history, strongly suggesting a Mendelian form of the disease, are good candidates for NGS-based studies.
Although GWAS have successfully revealed numerous susceptibility genes for common neurologic diseases, the odds ratios associated with these risk alleles are relatively low and account for only a small part of the estimated heritability, implicating that there are rare (present in <5% of the population), but not causative variants between these two extremes of the frequency spectrum, which usually have large effect size and cannot be captured by GWAS employing common SNPs (3). As a result of the development of NGS technologies, whole-exome sequencing (WES), or even whole-genome sequencing (WGS), have become considerably faster and more affordable over the past 5 years (4). In contrast to first-generation sequencing, also called Sanger sequencing, which can take several years and cost millions of dollars to sequence an entire diploid human genome, an NGS platform can produce the same genome sequence within a few weeks for as little as US$4,000-5,000 (5). More importantly, NGS technology have enabled the identification of rare variant with large effect size, including unmasking missense or nonsense single-base substitutions, as well as small insertions or deletions, which have important implications in risk prediction, diagnose, and treatment of neurological diseases (Figure 1 shows simplified workflows for NGS) (6-8).
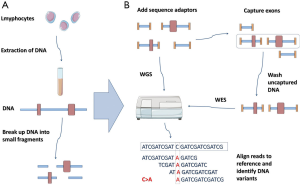
The value of NGS has been also demonstrated in the publishing explosion seen in the last three years, as more than 600 published studies used NGS to identify genes associated with a variety of disorders (5). In this article, we review recent NGS-based studies that aimed to investigate genetic causes for neurological diseases, including AD, PD, epilepsy, MS, stroke, ALS and spinocerebellar ataxias (SCA) (Table 1). In addition, the future directions of NGS applications are also discussed here.
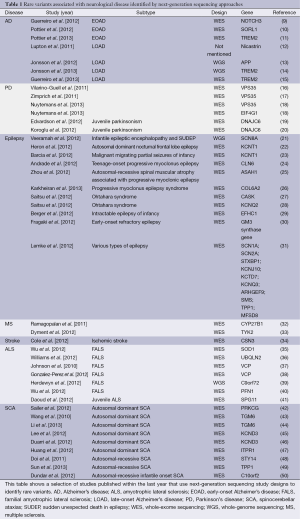
Full table
NGS approaches for genetic mapping of AD
Genetic factors are found to play a decisive role in the pathogenesis of AD (51). The mutations in APP, PSEN1 and PSEN2 are inherited in a Mendelian fashion, and directly lead to the early-onset AD (EOAD). However, numerous EOAD cases do not show any mutation in APP, PSEN1, or PSEN2, and this implicates that there are additional genetic factors involved in the pathogenesis of EOAD. In recent years, NGS has been applied to uncover genetic factors in small families with unexplained EOAD. To date, three NGS-based studies of people with EOAD have been published. The first study by Guerreiro et al. identified a missense mutation in NOTCH3 (p.R1231C), a gene previously linked to cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (9). The second WES study was conducted by Pottier and colleagues in patients with EOAD and identified missense and nonsense mutations in SORL1, which encodes a neuronal sorting protein that binds APP and directs it towards the endosome-recycling pathways (10). TREM2 is an immune phagocytic receptor expressed in brain microglia and modulates microglial phagocytosis and inflammatory pathway (52). More recently, by using WES, Pottier et al. investigated the association between TREM2 exon 2 variants and early-onset AD in a sample of Caucasian subjects of French origin. They found that the rs75932628-T variant (p.R47H) conferred a significant risk for EOAD (OR, 4.05; 95% CI, 1.3 to 16.8; P=0.009), and this variant was also revealed to be associated with the risk of late-onset AD (LOAD) (discussed in more detail below) (11).
In contrast, LOAD has been identified to be a complex disease which involves multiple susceptibility genes. Although GWAS have recently identified several risk variants for late-onset AD (53,54), these variants are generally associated with very low risk except the ε4 allele of apolipoprotein E, BIN 1 and CLU (55-57). Recently advances in NGS overcome the shortcomings of GWAS and add weight to the hypothesis that rare variation explains some of genetic heritability in AD. By using NGS, Lupton et al. investigated the effect of rare variants in the coding regions of the Nicastrin gene in a cohort of AD patients and identified a non-synonymous rare SNP (p.N417Y) with a statistically higher frequency in cases compared to controls in the Greek population (OR, 3.994; 95% CI, 1.105-14.439; P=0.035). Nicastrin is an obligatory component of the γ-secretase, the enzyme complex that leads to the production of Aβ fragments critically central to the pathogenesis of LOAD (12). Notably, by adopting NGS, two groups of researchers have uncovered the association between rare variants in TREM2 and LOAD. Jonsson et al. obtained whole genome sequence from a cohort of Icelanders, and found rs75932628-T variant in TREM2 that conferred a significant, three-fold increase in risk for AD. They then replicated the findings in cohorts from the USA, Norway, the Netherlands and Germany (14). Coincidentally, the same rare variant, rs75932628 was identified as causing a five-fold increase in risk for AD by Guerreiro and colleagues through WES technology. In addition, six other TREM2 variants were found in AD cases and not in controls, which may contribute to the pathogenesis of LOAD (15). Interestingly, through studying coding variants in APP in a set of whole-genome sequence data from 1,795 Icelanders, Jonsson and colleagues identified a protective APP missense variation (p.A676T) that protects against cognitive decline in the elderly without AD (13). This substitution is adjacent to the aspartyl protease β-site in APP, and results in an approximately 40% reduction in the formation of amyloidogenic peptides in vitro, supporting the notion that reduction of BACE1 cleavage protects against disease progression.
NGS approaches for genetic mapping of PD
In recent years, the understanding of the genetics of PD has seen great improvements: five genes are now known to be causative for monogenic forms (58-62), while 11 loci were recently identified as modulating risk for the development of common forms of PD (63). Nonetheless, a significant proportion of inherited PD cases still remain unexplained genetically. Recently, two independent studies utilized WES in an Austrian and a Swiss kindred, and identified the same p.D620N (c.1858G>A) mutation in the vacuolar protein sorting 35 homolog (VPS35) gene as the cause of autosomal dominant PD in these families (16,17). VPS35 is a component of the retromer complex and is involved in retrograde transport from the endosomes to the trans-Golgi network. The p.D620N mutation may lead to impaired recycling of membrane-associated proteins and dysfunctional endosomal-lysosomal trafficking. More recently, a NGS study conducted in 213 patients with PD and revealed 3 novel VPS35 variations (p.P316S; p.Y507F; p.E787K) that changed the coded amino acid which may contributed to the pathogenesis of PD (18). Additionally, a specific mutation in EIF4G1 (p.R1205H) was also identified as a strong PD risk factor in the same study (18).
Compared with late-onset disease, young-onset disease is more suitable for NGS-based studies, since young-onset disease is more likely due to multiple rare variants. By using homozygosity mapping and WES, Edvardson and colleagues identified a deleterious mutation in DNAJC6 (c.801-2A>G) in two patients with juvenile Parkinsonism. The mutation was associated with abnormal transcripts and marked reduced DNAJC6 mRNA level (19). Coincidentally, by mapping the disease locus with a lod score of 5.13 to a <3.5 Mbp region at 1p31.3 in a consanguineous family and subsequent WES analysis, Kӧroğlu et al. identified homozygous truncating mutation (p.Q734X) in DNAJC6. This findings further establish DNAJC6 as a juvenile parkinsonism gene, and expand the spectrums of the parkinsonism phenotype and DNAJC6 mutation (20).
The use of NGS in epilepsy
Epilepsies have a highly heterogeneous background with a strong genetic contribution. Mounting evidence has suggested that part of epilepsies are channelopathies, which caused by pathogenic variants in ion channel genes (64). Veeramah and colleagues performed WGS on a family containing an individual with infantile epileptic encephalopathy and sudden unexpected death in epilepsy, and discovered a de novo heterozygous missense mutation (c.5302A > G; p.N1768D) in the voltage-gated sodium-channel gene SCN8A. This mutation alters an evolutionarily conserved residue in Na (v) 1.6, one of the most abundant sodium channels in the brain. Analysis of the biophysical properties of the mutant channel demonstrated a dramatic increase in persistent sodium current, incomplete channel inactivation, and a depolarizing shift in the voltage dependence of steady-state fast inactivation (21). Meanwhile, mutations in KCNT1, encoding a sodium-gated potassium channel subunit, was recently found to cause the severe form of autosomal dominant nocturnal frontal lobe epilepsy (ADNFLE). Mutations were identified in three families by using WES, including c.2782C > T (p.R928C), c.2386T > C (p.Y796H) and c.1193G > A (p.R398Q). In addition, they also identified a de novo mutation in KCNT1, c.2688G > A (p.M896I), in a sporadic case with severe ADNFLE (22). Coincidentally, within the same gene, another four mutations (c.1283G > A, p.R428Q; c.2800G > A, p.A934T; c.1421G > A, p.R474H; c.2280C > G, p.I760M) that affected a highly conserved amino-acid residue were identified in patient with malignant migrating partial seizures of infancy. Functional studies showed that the mutations led to constitutive activation of the channel, mimicking the effects of phosphorylation of the C-terminal domain by protein kinase C. These results implicated the important role of ion channel complex in the pathogenesis of epilepsies (23).
Progressive myoclonic epilepsy (PME) represents a heterogeneous group of epilepsies characterized by myoclonic and generalized seizures with progressive neurological deterioration. Recently, mutations in three different genes have been identified as the genetic causes of this disease. Andrade et al. performed WES combined with homozygosity mapping in a multiplex family with autosomal recessive teenage-onset progressive myoclonus epilepsy and discovered a mutation (c.768C>G; p. D256E) in CLN6 which segregated with this disease (24). Besides, Zhou and colleagues identified a homozygous missense mutation (c.125C>T; p.T42M) in exon 2 of ASAH1 in the affected members of two unrelated PME families (25). More recently, Karkheiran et al. identified a homozygous, disease-segregating COL6A2 mutation (p.D215N) by adopting WES in two affected siblings in a consanguineous family with PME, and the authors suggested that that the COL6A2 p.D215N mutation is likely to be responsible for PME in this family, and additional studies are warranted to further establish the pathogenic role of COL6A2 mutations in the pathogenesis of PME (26).
Ohtahara syndrome (OS) is one of the epileptic encephalopathies characterized by severe and early-onset epilepsy, and several genes have been found to be associated with OS. As the development of NGS, new genes involved in the pathogenesis of OS have been uncovered recently. In a NGS-based study conducted by Saitsu et al., a 111-kb deletion involving exon 2 of CASK at Xp11.4 and a c.1A>G mutation in CASK gene have been identified. The detected mutations are highly likely to cause the loss of function of the CASK protein in affected individuals (27). In addition, by WES analysis of 12 patients with OS, these authors found 3 missense mutations in KCNQ2 (c.341C>T, p.T114I; c.1010C>G, p.A337G; c.794C>T, p.A265V) in 3 patients diagnosed with OS (28).
Almost one-third of patients with epilepsy cannot achieve seizure freedom and have a poor prognosis despite receiving adequate medical treatment and the administration of multiple anti-epileptic drugs (65). Although the mechanism remains unclear, the contributions of gene factors should not be neglected for this refractoriness. In a nonconsanguineous Moroccan-Jewish family, where three of their seven children presented with refractory epilepsy, Berger and colleagues performed WES and revealed a pathogenic mutation (p.F229L) in the EFHC1 gene that cosegregated with this disease (29). Besides, Fragaki et al. uncovered a homozygous loss-of-function mutation in exon 6 of the GM3 synthase gene (c.862C>T; p.R288*), which caused the deficiency of GM3 synthase and was responsible for an early-onset refractory epilepsy (30).
Notably, Lemke and colleagues recently performed NGS of 265 channel genes, which containing the most relevant epilepsy genes and covering the most relevant epilepsy phenotypes known so far and detected multiple pathogenic mutations in 10 genes (SCN1A: p.R222X, p. E289V, p.379R, p.R393H; SCN2A: p.V208E; STXBP1: p.R122X; KCNJ10: p.L68P, p.I129V; KCTD7: p.L108M; KCNQ3: p.P574S; ARHGEF9: p.R290H; SMS: p.F58L; TPP1: p.Q278R, p.Q422H; MFSD8: p.T294K). All these mutations have been confirmed by conventional Sanger sequencing and validated by parental testing and segregation analysis. Together with the studies discussed above, this study further proved NGS is a powerful diagnostic tool that contribute to collecting information on both common and unknown epileptic disorders and in delineating associated phenotypes of less frequently mutated genes (31).
The use of NGS in MS
Although the cause of MS still remains elusive, the genetic factors that contribute to the susceptibility of MS are undeniable. The HLA class II as well as non-HLA loci that associated with this disease have been identified by GWAS (66). However, these alleles are neither necessary nor sufficient to cause MS, and rare variants that could have larger effect sizes may be neglected by GWAS. To date, two rare variants contributing to MS susceptibility have identified by WES. Ramagopalan and colleagues performed WES in a cohort of MS patients and identified a rare functional variant (p.R389H) within CYP27B1, the gene encodes the vitamin D-activating 1-alpha hydroxylase enzyme. This variant leads to loss of gene function, which affects 1-α-hydroxylation of vitamin D and is associated with MS susceptibility (32). Besides, a novel mutation (rs55762744) in TYK2, a tyrosine kinase that modulates the function of multiple immune-related genes, was also identified in an extended pedigree with an unusually high frequency of MS by Dyment and colleagues (33).
The use of NGS in stroke
Epidemiological studies suggest that genetic risk factors are important for stroke, and due to the heterogeneity of the stroke phenotype and the difficulties in obtaining large sample sizes, GWAS in stroke have lagged behind those in other cerebrovascular diseases. Meanwhile, most genetic variants discovered using GWAS account for only a small increase in disease risk, with odds ratios most often between 1.1 and 1.3 (67). Considering the inherent shortage of GWAS in detecting low frequency variants, NGS is now gaining more attention in identification of these variants, especially in young-onset disease, since young-onset disease is more likely due to multiple rare variants as compared to late-onset disease. In 2012, Cole and colleagues launched a pilot study in 10 young-onset ischemic stroke cases including 8 African-Americans and 2 Caucasians, and identified a coding polymorphism within CSN3, a gene encoding kappa-casein was associated with young-onset ischemic stroke (34). Of note, this is the only NGS-based study that identified variants involved in the pathogenesis of stroke.
NGS to identify genetic causes of ALS
Of all ALS cases, approximately 10% are familial ALS (FALS), typically with a dominant inheritance mode (68). Four genes are relatively frequently encountered in familial ALS: SOD1, TDP-43, FUS and VCP. Mutations in SOD1 are responsible for 20% of familial ALS, and over 160 SOD1 pathogenic mutations have been identified so far. Recently, Wu and colleagues performed WES in a Chinese pedigree and identified a novel mutation (Cys146X) in SOD 1 gene as a causative factor in FALS (35). In addition to SOD1, mutations in VCP are responsible for approximately 1.5% of FALS cases. By using WES, Johnson and colleagues identified a novel mutation (p.R191Q) in the VCP gene in an Italian family with autosomal dominantly inherited ALS (37). The same mutation in VCP was also found in a 4-generation Israeli-Arab family with atypical ALS by González-Pérez et al., further supporting the view that motor neuron disease is part of the clinical spectrum of VCP-associated disease (38).
However, in addition to FALS caused by mutations in above-mentioned genes, roughly 50% of ALS families remain unexplained after routine genetic testing. Recent NGS-based studies have identified several genetic causes which provide explanation to these cases. By performing WGS in five family members from a pedigree with autosomal dominant classical ALS, Herdewyn and colleagues identified a hexanucleotide repeat expansion in the noncoding region of C9orf72 as the cause of chromosome 9-linked ALS (39). Besides, WES of two large ALS families has identified 4 mutations (p.C71G, p.M114T, p.E117G, p.G118V) within the PFN1 gene that can cause FALS (40). More recently, Williams et al. adopted WES and found a novel missense UBQLN2 mutation (c.1460C>T, p.T487I) in 2 apparently unrelated multigenerational ALS families, further support the importance of ubiquilin 2 in the pathogenesis of ALS (36). In addition to adult-onset ALS, Daoud and colleagues identified a mutation (c.5199delA) in exon 30 of SPG11 gene that was associated with atypical juvenile ALS by performing WES (41).
NGS to identify genetic causes of SCA
Autosomal dominant SCA constitute a large, heterogeneous group of progressive neurodegenerative diseases with multiple types. To date, 32 dominant SCAs have been chromosomally mapped, and the genes causing 20 of these disorders have so far been identified (69). The genetic etiologies of many autosomal dominant SCAs have yet to be elucidated, partly due to the limitations of traditional positional cloning strategies in finding causative genes of rare Mendelian disorders. In recent years, NGS-based studies open up new avenues of identification of causative genes in autosomal dominant SCA. By using WES, Sailer et al. investigated a large, 5-generational British kindred with autosomal dominant SCA and identified a novel pathogenic mutation (p.R26G) in the PRKCG gene. Furthermore, this variant was confirmed using Sanger sequencing and showed segregation with disease in the entire family (42). Recently, Wang et al. sequenced the whole exome of four patients in a Chinese four-generation spinocerebellar ataxia family and identified a novel missense mutation, c.1550T>G transition (p.L517W), in exon 10 of TGM6. This mutation is at a highly conserved position, and is predicted to have a functional impact, and completely cosegregated with the phenotype. These authors also further extend their finding by identifying another missense mutation c.980A>G transition (p.D327G) in exon 7 of TGM6 in an additional SCA family, which also cosegregated with the phenotype (43). Within the same gene, Li and colleagues performed WES in a three-generation Chinese family with SCA and found a novel mutation (c.1528G>C), which showed perfect co-segregation with disease phenotype in all nine members of this family. This result further confirmed previous finding from Wang et al. that mutations in TGM6 gene represent an important cause of SCA in Chinese (44).
Ion channels functions as a key regulators of neuronal excitability in the pathogenesis of SCA. In SCA22 patients of diverse ethnic origins, Lee et al. performed WES and identified an in-frame 3-nucleotide deletion (c.679_681delTTC p.F227del) and 3 mutations (c.1034G>T, p.G345V; c.1013T>C, p.V338E and c.1130C>T, p.T377M) in the voltage-gated potassium channel K(v)4.3-encoding gene KCND3, which was associated with this disease (45). At the same time, another mutation (p.T352P) within the same gene has been identified in a large SCA19 family by Duarri and colleagues through WES. These authors suggested that KCND3 mutations may cause SCA by impairing protein maturation and/or reducing channel function (46). ITPR1 encodes inositol 1,4,5-trisphosphate receptor, type-1, a ligand-gated ion channel that mediates calcium release from the endoplasmic reticulum. In addition to SCA15, alteration of ITPR1 function can cause a distinct congenital nonprogressive ataxia, as demonstrated by a recent study from Huang and colleagues. They performed WES in a Australian family and a Canadian family and identified two pathogenic missense mutations (c.4657G >A, p.V1553M; c.1804A >G, p.N602D), highlighting a significant role of the ITPR1-related pathway in the development and maintenance of the normal functions of the cerebellum (47).
In addition to autosomal dominant forms of SCA, NGS also offers a very powerful method to discover the genetic factors of other types of SCA. Autosomal recessive SCA (ARSCAs) are clinically and genetically heterogeneous disorders associated with diverse neurological and nonneurological features that occur before the age of 20. Using WES combined with homozygosity mapping, Doi and colleagues identified a homozygous missense mutation in SYT14 (p.G484D), encoding synaptotagmin XIV (SYT14), in a Japanese family in which two siblings have slow progression of a type of ARSCA with psychomotor retardation. These authors suggested that this mutation within SYT14 lead to the alteration of the membrane-trafficking machinery, which may represent a distinct pathomechanism associated with ARSCAs (48). In addition, by using WES, a missense (c.1397T>G, p.V466G) and a splice site variant (c.509-1G>C) in TPP1, cosegregating with the disease, were found in a previously described ARSCA family as well as in another patient with an ARSCA phenotype by Sun and colleagues, suggesting a pathogenic role of TPP1 mutations in ARSCA (49). Dundar and colleagues applied WES to a family containing two individuals with infantile onset spinocerebellar ataxia (IOSCA), and identified a novel homozygous missense mutation c.1366C>G (p.L456V) in C10orf2, which may contributed to the pathogenesis of IOSCA (50).
Conclusions and future perspectives
Although GWAS have made great improvement in understanding the genetic causes of complex neurological diseases, a large proportion of the genetic etiology of these disorders remains unexplained. NGS platforms could circumvent a large number of limitations of GWAS, and offer powerful new tools to dissect out genetic contributions to disease etiology. As discussed above, WES and WGS have identified numerous rare variants important in complex neurological diseases, as well as Mendelian neurological conditions. Currently, WES has wider applications than WGS, largely because of its relatively lower cost (the exome is approximately 1% of the whole genome) and the notion that most sequence variations leading to a severe phenotypic effect are located in the coding part of the genome. However, increasing evidence suggests that non-coding variants cause or increase the risk of neurodegenerative disease. Since WGS provides better coverage than WES, and the cost of sequencing continues to drop, it will eventually be more time-efficient and cost-effective to perform the WES in patients rather than to sequence the exome only. In addition to WES and WGS, functional genomics methods, such as RNA-seq and ChIP-seq, may open up new ways of identifying disease-associated biochemical pathways. The combinatorial method to uncover the genetic basis of complex neurological disease will lead to huge advances in our understanding, with applications for diagnostics and potentially novel treatments.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81000544, 81171209), the Shandong Provincial Natural Science Foundation, China (ZR2010HQ004, ZR2011HZ001), the Medicine and Health Science Technology Development Project of Shandong Province (2011WSA02018, 2011WSA02020), and the Shandong Provincial Outstanding Medical Academic Professional Program.
Disclosure: The authors declare no conflict of interest.
References
- Urdinguio RG, Sanchez-Mut JV, Esteller M. Epigenetic mechanisms in neurological diseases: genes, syndromes, and therapies. Lancet Neurol 2009;8:1056-72. [PubMed]
- Simón-Sánchez J, Singleton A. Genome-wide association studies in neurological disorders. Lancet Neurol 2008;7:1067-72. [PubMed]
- Pritchard JK. Are rare variants responsible for susceptibility to complex diseases? Am J Hum Genet 2001;69:124-37. [PubMed]
- Metzker ML. Sequencing technologies - the next generation. Nat Rev Genet 2010;11:31-46. [PubMed]
- Thompson R, Drew CJ, Thomas RH. Next generation sequencing in the clinical domain: clinical advantages, practical, and ethical challenges. Adv Protein Chem Struct Biol 2012;89:27-63. [PubMed]
- Handel AE, Disanto G, Ramagopalan SV. Next-generation sequencing in understanding complex neurological disease. Expert Rev Neurother 2013;13:215-27. [PubMed]
- Foo JN, Liu JJ, Tan EK. Whole-genome and whole-exome sequencing in neurological diseases. Nat Rev Neurol 2012;8:508-17. [PubMed]
- Bras J, Guerreiro R, Hardy J. Use of next-generation sequencing and other whole-genome strategies to dissect neurological disease. Nat Rev Neurosci 2012;13:453-64. [PubMed]
- Guerreiro RJ, Lohmann E, Kinsella E, et al. Exome sequencing reveals an unexpected genetic cause of disease: NOTCH3 mutation in a Turkish family with Alzheimer's disease. Neurobiol Aging 2012;33:1008.e17-23.
- Pottier C, Hannequin D, Coutant S, et al. High frequency of potentially pathogenic SORL1 mutations in autosomal dominant early-onset Alzheimer disease. Mol Psychiatry 2012;17:875-9. [PubMed]
- Pottier C, Wallon D, Rousseau S, et al. TREM2 R47H variant as a risk factor for early-onset alzheimer’s disease. J Alzheimers Dis 2013;35:45-9. [PubMed]
- Lupton MK, Proitsi P, Danillidou M, et al. Deep sequencing of the Nicastrin gene in pooled DNA, the identification of genetic variants that affect risk of Alzheimer’s disease. PLoS One 2011;6:e17298. [PubMed]
- Jonsson T, Atwal JK, Steinberg S, et al. A mutation in APP protects against Alzheimer’s disease and age-related cognitive decline. Nature 2012;488:96-9. [PubMed]
- Jonsson T, Stefansson H, Steinberg S, et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N Engl J Med 2013;368:107-16. [PubMed]
- Guerreiro R, Wojtas A, Bras J, et al. TREM2 variants in Alzheimer’s disease. N Engl J Med 2013;368:117-27. [PubMed]
- Vilariño-Güell C, Wider C, Ross OA, et al. VPS35 mutations in Parkinson disease. Am J Hum Genet 2011;89:162-7. [PubMed]
- Zimprich A, Benet-Pagès A, Struhal W, et al. A mutation in VPS35, encoding a subunit of the retromer complex, causes late-onset Parkinson disease. Am J Hum Genet 2011;89:168-75. [PubMed]
- Nuytemans K, Bademci G, Inchausti V, et al. Whole exome sequencing of rare variants in EIF4G1 and VPS35 in Parkinson disease. Neurology 2013;80:982-9. [PubMed]
- Edvardson S, Cinnamon Y, Ta-Shma A, et al. A deleterious mutation in DNAJC6 encoding the neuronal-specific clathrin-uncoating co-chaperone auxilin, is associated with juvenile parkinsonism. PLoS One 2012;7:e36458. [PubMed]
- Köroğlu Ç, Baysal L, Cetinkaya M, et al. DNAJC6 is responsible for juvenile parkinsonism with phenotypic variability. Parkinsonism Relat Disord 2013;19:320-4. [PubMed]
- Veeramah KR, O’Brien JE, Meisler MH, et al. De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP. Am J Hum Genet 2012;90:502-10. [PubMed]
- Heron SE, Smith KR, Bahlo M, et al. Missense mutations in the sodium-gated potassium channel gene KCNT1 cause severe autosomal dominant nocturnal frontal lobe epilepsy. Nat Genet 2012;44:1188-90. [PubMed]
- Barcia G, Fleming MR, Deligniere A, et al. De novo gain-of-function KCNT1 channel mutations cause malignant migrating partial seizures of infancy. Nat Genet 2012;44:1255-9. [PubMed]
- Andrade DM, Paton T, Turnbull J, et al. Mutation of the CLN6 gene in teenage-onset progressive myoclonus epilepsy. Pediatr Neurol 2012;47:205-8. [PubMed]
- Zhou J, Tawk M, Tiziano FD, et al. Spinal muscular atrophy associated with progressive myoclonic epilepsy is caused by mutations in ASAH1. Am J Hum Genet 2012;91:5-14. [PubMed]
- Karkheiran S, Krebs CE, Makarov V, et al. Identification of COL6A2 mutations in progressive myoclonus epilepsy syndrome. Hum Genet 2013;132:275-83. [PubMed]
- Saitsu H, Kato M, Osaka H, et al. CASK aberrations in male patients with Ohtahara syndrome and cerebellar hypoplasia. Epilepsia 2012;53:1441-9. [PubMed]
- Saitsu H, Kato M, Koide A, et al. Whole exome sequencing identifies KCNQ2 mutations in Ohtahara syndrome. Ann Neurol 2012;72:298-300. [PubMed]
- Berger I, Dor T, Halvardson J, et al. Intractable epilepsy of infancy due to homozygous mutation in the EFHC1 gene. Epilepsia 2012;53:1436-40. [PubMed]
- Fragaki K, Ait-El-Mkadem S, Chaussenot A, et al. Refractory epilepsy and mitochondrial dysfunction due to GM3 synthase deficiency. Eur J Hum Genet 2013;21:528-34. [PubMed]
- Lemke JR, Riesch E, Scheurenbrand T, et al. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia 2012;53:1387-98. [PubMed]
- Ramagopalan SV, Dyment DA, Cader MZ, et al. Rare variants in the CYP27B1 gene are associated with multiple sclerosis. Ann Neurol 2011;70:881-6. [PubMed]
- Dyment DA, Cader MZ, Chao MJ, et al. Exome sequencing identifies a novel multiple sclerosis susceptibility variant in the TYK2 gene. Neurology 2012;79:406-11. [PubMed]
- Cole JW, Stine OC, Liu X, et al. Rare variants in ischemic stroke: an exome pilot study. PLoS One 2012;7:e35591. [PubMed]
- Wu J, Shen E, Shi D, et al. Identification of a novel Cys146X mutation of SOD1 in familial amyotrophic lateral sclerosis by whole-exome sequencing. Genet Med 2012;14:823-6. [PubMed]
- Williams KL, Warraich ST, Yang S, et al. UBQLN2/ubiquilin 2 mutation and pathology in familial amyotrophic lateral sclerosis. Neurobiol Aging 2012;33:2527.e3-10.
- Johnson JO, Mandrioli J, Benatar M, et al. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 2010;68:857-64. [PubMed]
- González-Pérez P, Cirulli ET, Drory VE, et al. Novel mutation in VCP gene causes atypical amyotrophic lateral sclerosis. Neurology 2012;79:2201-8. [PubMed]
- Herdewyn S, Zhao H, Moisse M, et al. Whole-genome sequencing reveals a coding non-pathogenic variant tagging a non-coding pathogenic hexanucleotide repeat expansion in C9orf72 as cause of amyotrophic lateral sclerosis. Hum Mol Genet 2012;21:2412-9. [PubMed]
- Wu CH, Fallini C, Ticozzi N, et al. Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis. Nature 2012;488:499-503. [PubMed]
- Daoud H, Zhou S, Noreau A, et al. Exome sequencing reveals SPG11 mutations causing juvenile ALS. Neurobiol Aging 2012;33:839.e5-9.
- Sailer A, Scholz SW, Gibbs JR, et al. Exome sequencing in an SCA14 family demonstrates its utility in diagnosing heterogeneous diseases. Neurology 2012;79:127-31. [PubMed]
- Wang JL, Yang X, Xia K, et al. TGM6 identified as a novel causative gene of spinocerebellar ataxias using exome sequencing. Brain 2010;133:3510-8. [PubMed]
- Li M, Pang S, Song Y, et al. Whole exome sequencing identifies a novel mutation in the transglutaminase 6 gene for spinocerebellar ataxia in a Chinese family. Clin Genet 2013;83:269-73. [PubMed]
- Lee YC, Durr A, Majczenko K, et al. Mutations in KCND3 cause spinocerebellar ataxia type 22. Ann Neurol 2012;72:859-69. [PubMed]
- Duarri A, Jezierska J, Fokkens M, et al. Mutations in potassium channel kcnd3 cause spinocerebellar ataxia type 19. Ann Neurol 2012;72:870-80. [PubMed]
- Huang L, Chardon JW, Carter MT, et al. Missense mutations in ITPR1 cause autosomal dominant congenital nonprogressive spinocerebellar ataxia. Orphanet J Rare Dis 2012;7:67. [PubMed]
- Doi H, Yoshida K, Yasuda T, et al. Exome sequencing reveals a homozygous SYT14 mutation in adult-onset, autosomal-recessive spinocerebellar ataxia with psychomotor retardation. Am J Hum Genet 2011;89:320-7. [PubMed]
- Sun Y, Almomani R, Breedveld GJ, et al. Autosomal recessive spinocerebellar ataxia 7 (SCAR7) is caused by variants in TPP1, the gene involved in classic late-infantile neuronal ceroid lipofuscinosis 2 disease (CLN2 Disease). Hum Mutat 2013;34:706-13. [PubMed]
- Dündar H, Ozgül RK, Yalnızoğlu D, et al. Identification of a novel Twinkle mutation in a family with infantile onset spinocerebellar ataxia by whole exome sequencing. Pediatr Neurol 2012;46:172-7. [PubMed]
- Hollingworth P, Harold D, Jones L, et al. Alzheimer's disease genetics: current knowledge and future challenges. Int J Geriatr Psychiatry 2011;26:793-802. [PubMed]
- Jiang T, Yu JT, Zhu XC, et al. TREM2 in Alzheimer's disease. Mol Neurobiol 2013;48:180-5. [PubMed]
- Harold D, Abraham R, Hollingworth P, et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genet 2009;41:1088-93. [PubMed]
- Hollingworth P, Harold D, Sims R, et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nat Genet 2011;43:429-35. [PubMed]
- Bu G. Apolipoprotein E and its receptors in Alzheimer’s disease: pathways, pathogenesis and therapy. Nat Rev Neurosci 2009;10:333-44. [PubMed]
- Hu X, Pickering E, Liu YC, et al. Meta-analysis for genome-wide association study identifies multiple variants at the BIN1 locus associated with late-onset Alzheimer's disease. PLoS One 2011;6:e16616. [PubMed]
- Yu JT, Tan L. The role of clusterin in Alzheimer’s disease: pathways, pathogenesis, and therapy. Mol Neurobiol 2012;45:314-26. [PubMed]
- Valente EM, Abou-Sleiman PM, Caputo V, et al. Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 2004;304:1158-60. [PubMed]
- Paisán-Ruíz C, Jain S, Evans EW, et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 2004;44:595-600. [PubMed]
- Bonifati V, Rizzu P, van Baren MJ, et al. Mutations in the DJ-1 gene associated with autosomal recessive early-onset parkinsonism. Science 2003;299:256-9. [PubMed]
- Kitada T, Asakawa S, Hattori N, et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 1998;392:605-8. [PubMed]
- Polymeropoulos MH, Lavedan C, Leroy E, et al. Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 1997;276:2045-7. [PubMed]
- International Parkinson Disease Genomics Consortium, Nalls MA, Plagnol V, et al. Imputation of sequence variants for identification of genetic risks for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet 2011;377:641-9. [PubMed]
- Jurkat-Rott K, Lerche H, Weber Y, et al. Hereditary channelopathies in neurology. Adv Exp Med Biol 2010;686:305-34. [PubMed]
- Callaghan BC, Anand K, Hesdorffer D, et al. Likelihood of seizure remission in an adult population with refractory epilepsy. Ann Neurol 2007;62:382-9. [PubMed]
- International Multiple Sclerosis Genetics Consortium, Hafler DA, Compston A, et al. Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med 2007;357:851-62. [PubMed]
- Markus HS. Stroke genetics: prospects for personalized medicine. BMC Med 2012;10:113. [PubMed]
- Dion PA, Daoud H, Rouleau GA. Genetics of motor neuron disorders: new insights into pathogenic mechanisms. Nat Rev Genet 2009;10:769-82. [PubMed]
- Klockgether T. Update on degenerative ataxias. Curr Opin Neurol 2011;24:339-45. [PubMed]


