
A pilot study of expanded newborn screening for 573 genes related to severe inherited disorders in China: results from 1,127 newborns
Introduction
Newborn screening (NBS), a successful public health program that prevents morbidity and mortality through early detection and management of specific conditions, can improve the life expectation of children with certain life-threatening metabolic disorders by allowing timely therapeutic interventions. The screening of newborns for inherited metabolic diseases (IMDs) and other common genetic disorders is now a routine component of neonatal care in many countries now. Tandem mass spectrometry (MS/MS) is widely used to detect metabolites related to amino acid metabolism, organic acid metabolism and fatty acid oxidation in dried blood spots (DBS), and is currently the principal method used in NBS centers to detect biochemical metabolites in developed countries (1). In China, apart from MS/MS screening, biochemical NBS also includes phenylalanine (Phe) test for hyperphenylalaninemia, thyroid-stimulating hormone (TSH) test for hyperthyroidism or hypothyroidism, 17-α-hydroxyprogesterone (17-OHP) test for congenital adrenal hyperplasia (CAH), and glucose-6-phosphate dehydrogenase (G6PD) enzymatic analysis for G6PD deficiency.
Next-generation sequencing (NGS) technologies has enabled high-throughput detection of genetic variants. The rapid development of NGS has reduced the cost of sequencing and genetic analysis such that NGS can now be used in NBS. Genome-wide sequencing in the US has been discussed and been applied in 159 newborns (2,3). The results suggested that newborn genomic sequencing can effectively detect the risk and carrier status for a wide range of disorders that are not detectable by current NBS assays (4). Genomic sequencing of newborns can help avoid diagnostic odysseys in ill newborns, allow optimal early treatment strategies for affected newborns (5,6), and identify carrier status that could help reproductive planning in future. Despite the anticipated benefits, the major challenges hindering its application include data interpretation and appropriate reporting of information. In our previous research, we designed a panel of 77 genes related to over 40 IMDs and demonstrated that genomic DNA extracted from DBS could generate reliable sequencing data, which would enable NGS to function as a second-tier diagnostic test in NBS (7).
Considering these benefits and challenges, we launched a project to screen newborns for gene variants related to severe inherited disorders. 573 genes were selected based on prevalence and age of onset of the diseases. 1,173 newborns born in 2016 in the NBS center of Xinhua Hospital in Shanghai were randomly enrolled in this study. They had been screened for biochemical NBS and had went through NGS. 1,127 out of 1,173 samples returned qualified NGS data. We compared the biochemical results with genetic variants in these newborns and attempt to investigate whether NGS could identify neonates with severe inherited disorders that were confirmed by current biochemical screening effectively and whether NGS screening could be used as a supplement to improve the detection rate of biochemical screening.
Previous studies have demonstrated that the frequencies of Mendelian disorders in China vary according to the diverse ethnic backgrounds and areas (8). Most of the newborns recruited in this study were born in Shanghai and belonged to Han population, which is the largest ethnic group in China. IMDs, a group of rare diseases with an overall incidence of 1/500 in all ethnic cohorts, are caused by enzyme or transporter abnormalities in biochemical pathways for metabolism (9). IMDs are genetically heterogeneous disorders and are mostly inherited in autosomal recessive inheritance pattern. Patients with IMDs may present severe symptoms. For disorders related to amino acid metabolism, organic acid metabolism, and fatty acid oxidation tested by MS/MS, the diagnostic rate was between 1:10,165 to 1:2,363 according to previous NBS studies of large cohorts in three provinces of China (10-12). This study also aimed to investigate the carrier frequencies of mutations in genes related to amino acid metabolism, organic acid metabolism, and fatty acid oxidation disorders in this cohort.
We present the following article in accordance with the STROBE reporting checklist (available at http://dx.doi.org/10.21037/atm-20-1147).
Methods
Subjects
This study included 1,173 residual DBS specimens from newborns (600 males and 573 females) at Xinhua Hospital in Shanghai. All the newborns were randomly selected in September 2016. 46 DBS failed to meet quality control standards. DNA extracted from 1,127 samples (576 males and 551 females) was subjected to sequencing. Among these newborns, 1052 were born in Shanghai (93.35%, 535 males and 517 females) and 75 were born in Macao (6.65%, 42 males and 33 females). All the newborns had undergone biochemical NBS, including MS/MS, Phe, TSH, 17-OHP, and G6PD analysis. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and was approved by the Xinhua Hospital Ethics Committee Affiliated to Shanghai Jiao Tong University School of Medicine (XHEC-C-2017-021-2). Written consent was exempted as the residual DBS samples were reused in this study.
Target diseases and genes
The causative genes for severe inherited disorders included hematological, cardiovascular, immunological, gastrointestinal, endocrinological, skeletal, neurological, muscular, dermatological, urinary, respiratory, and IMDs. A total of 573 genes related to more than 200 birth-defect diseases were selected based on prevalence and age of onset (available on: https://cdn.amegroups.cn/static/application/8168bab76b1f94f75f011efab391f819/atm-20-1147-1.pdf). Rare diseases with extremely low incidence rates were excluded.
NGS and data analysis
Target-specific primers for amplicons were generated based on the hg19/GRCh37 human reference genome. The primers were designed to have similar length, GC content, and similar amplicon fragment size to maximize the amplification efficiency. The amplicon fragments included the exons and flanking intronic sequences. DNA extracted from a single DBS was generally enough for target capture and library preparation. The amplicons were sequenced on a NovaSeq 6000 instrument (Illumina, San Diego, CA, USA) according to the manufacturer’s protocol. Paired-end reads were aligned to the NCBI reference sequence (hg19/GRCh37) using BWA and variant calls were made using GATK. Primary analyses of the NGS data included sequence alignment, post-processing, and variant calling. The mean sequencing coverage was 305× and on average 99.4% of bases for every sample were sequenced to at least 20× coverage.
Variants met one of the following inclusion criteria were selected as candidate variants: (I) variants classified as pathogenic/likely pathogenic mutations in ClinVar database; (II) variants classified as disease-causing mutations (DM) or probable disease-causing mutations (DM?) in the Human Gene Mutation Database (HGMD); (III) frameshift, stop-gained, stop-lost, and splice donor/acceptor variants that were absent in ClinVar or HGMD. Considering there are false positive variants in HGMD, we set the criteria of upper frequency limit to reduce the false positive rate. For candidate variants that were absent in ClinVar database and only appeared in HGMD, the variants with >0.2% frequency in the population variant databases—Genome Aggregation Database (gnomAD), and 1000Genomes of East Asia database were filtered. Data from our NBS center revealed that hyperphenylalaninemia caused by PAH mutations is the most common IMDs except for G6PD deficiency that could only be triggered by exposure to exogenous primaquine or fava beans. Hence, we referred to the criteria for BS1 (0.2%) of PAH as the upper frequency limit for variants that only appeared in HGMD (13). Furthermore, in case two variants were identified in an autosomal recessive disorder that both met any piece of the three criteria in a subject, or one variant was identified in an autosomal dominant or X-linked disorder that met any one of the three criteria, the variants were exempted from the frequency limitation and further evaluated. Biochemical data were compared with the information on genetic variants. Newborns with candidate variants were followed up for further evaluation.
Results
Comparison of biochemical screening to genetic variants
In the biochemical screening, four infants (all males, ID: 83162, 83841, 83847, 84010) showed low G6PD enzymatic activity. Three newborns (ID: 83426, 83904, 84144) showed elevated 17-OHP levels at the initial screening, who turned out to be false positives in the followed confirmation test. No specimen gave abnormal result on TSH, Phe and MS/MS.
The four newborns carried hemizygous G6PD mutations that were classified as disease causing in HGMD. They are the same four babies with low G6PD enzyme levels, indicating the high sensitivity and specificity of G6PD deficiency test. Causative variants for CAH were not found in the cohort.
Apart from the four samples identified by biochemical screening, we found one individual (ID: 84123) carried two SLC22A5 variants and two individuals (ID: 84066, 84193) each carried two DUOX2 variants. Since the DUOX2 variants (c.1588A>T and c.1606C>T) of newborn 84193 could be phased in cis as the two variants are close, this case was excluded. The parents of individual 84066 were lost contact.
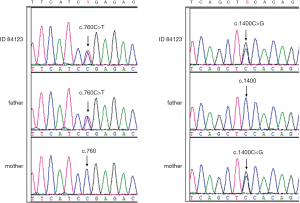
In 2019, we recalled the individual 84123 and verified the compound heterozygous variants of SLC22A5. The variant c.760C>T (p.R254X) was inherited from his father and c.1400C>G (p.S467C) was inherited from his mother (Figure 1). The two variants are common mutations in patients with carnitine deficiency(14). MS/MS screening results for individual 84123 in 2019 showed reduced free carnitine deficiency (C0 value: 4.3 µmol/L, reference: 10–60 µmol/L), which was consistent with the results of the genetic analysis, indicating a false-negative MS/MS finding at birth (C0 value: 11.6 µmol/L, reference: 10–60 µmol/L).
Individuals with candidate variants in IMDs related genes were listed in Table 1.

Full table
IMDs carrier frequencies
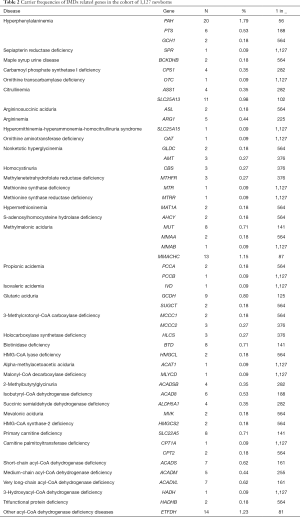
We calculated the carrier frequencies of mutations in 77 causative genes related to amino acid metabolism, organic acid metabolism, and fatty acid oxidation disorders that may be detected by MS/MS screening (available on: https://cdn.amegroups.cn/static/application/05176008dca3731212e9665e49807618/atm-20-1147-2.pdf). The top five genes with the highest carrier frequencies in these newborns were PAH (1:56, 1.79%), ETFDH (1:81, 1.23%), MMACHC (1:87, 1.15%), SLC25A13 (1:102, 0.98%), and GCDH (1:125, 0.80%) based on our criteria for clinically relevant variants. Carrier frequencies of IMDs related genes in this cohort was listed in Table 2. Candidate variants of the other 26 genes were not identified in the cohort: QDPR, PCBD1, BCKDHA, DBT, DLD, FAH, TAT, HPD, GCSH, HGD, MMADHC, MCEE, AUH, TAZ, OPA3, DNAJC19, ASPA, ETHE1, L2HGDH, D2HGDH, IDH2, SLC25A20, ETFA, ETFB, DECR1, and HADHA.
Full table
Discussion
Over the last decade, NGS methods have been proven to be effective for detecting genetic disorders (15). The cost of NGS has decreased greatly. Our previous study demonstrated that DNA extracted from DBS could be used for NGS (7). In this study, we proved that combining NGS screening with biochemical screening could improve the current NBS efficiency.
G6PD deficiency is an X-linked disorder due to G6PD mutations and is the most common human enzyme defect. The most frequent clinical manifestations of G6PD deficiency are neonatal jaundice and acute hemolytic anemia, which are usually triggered by an exogenous agent. Heterozygous females generally have less severe clinical manifestations compared to G6PD-deficient males (16). In this cohort, 535 males were born in Shanghai, two of whom (ID: 83162, 84010) showed G6PD deficiency. Among 42 males born in Macao, which is located in the subtropical area of Asia, two males (ID: 83841, 83847) showed G6PD deficiency. These findings indicated the higher prevalence of G6PD deficiency in Macao than that in Shanghai.
Three infants had high 17-OHP levels at birth, which turned out to be false positive when they were followed in a short time interval. Reexamination for neonates could be time-consuming. CAH included a group of disorders involving defects in cortisol synthesis, with the prevalence of about one in 10,000 births (17). According to previous studies, 17-OHP screening for CAH is facing challenges because of a high false-positive rate and a low positive predictive value (18-20). However, the NGS procedure for CAH is also technically challenging as the highly identical pseudogene of CYP21A2 makes it difficult to assess the accuracy of sequencing. Methods of better sensitivity and specificity for CAH screening are required.
DUOX2 is related to thyroid hormone synthesis dysfunction (21). One case (ID 84066) carried two DUOX2 variants, c.3391G>T (p.A1131S) and c.2202G>A (p.W734X). Whether the two DUOX2 variants were in trans could not be distinguished. Thus, samples from the parents were required for verification. Our review of the related literature revealed a report of a patient with congenital hypothyroidism carried compound heterozygous variations p.W734X and p.A1131S on DUOX2 (22). The patient (patient 1) in the literature showed a high TSH level. According to the guidelines for variant interpretation from ACMG/AMP, c.2202G>A(p.W734X) can be interpreted as a pathogenic variant (PVS1 + PM2 + PP4), and c.3391G>T(p.A1131S) can be interpreted as a likely pathogenic variant (PM1 + PM2 + PM3 + PP3 + PP4) (23). The newborn (ID: 84066) in this study carried the same two variants and. This infant, with normal TSH level in the initial screening (1.5 mU/L; the cut-off value for TSH in our lab is 10 mU/L) was born at full term (38.5 weeks) with normal weight (3.095 kg); therefore, the possibility of late-onset TSH increase was low. However, we failed to contact the family of the newborn for further study. We propose two possible explanations for these findings. First, the two variants might be in cis. Thus, there would be no impact on her thyroid function. Second, the biochemical result was false-negative. Since the parents are unable to contact, we could not draw valid conclusion of the case. This points out the proband-only sequencing approach made it hard to distinguish whether the two variants were in trans or in cis on the gene, except for the two variants were close enough to be phased by bioinformatic methods.
Biallelic mutations of SLC22A5 can cause systemic primary carnitine deficiency (SPCD) and lead to impaired fatty acid oxidation in muscles (24). SPCD encompasses a broad clinical spectrum including metabolic decompensation in infancy, childhood myopathy involving heart and skeletal muscle, pregnancy-related decreased stamina or exacerbation of cardiac arrhythmia, fatigability in adulthood, or absence of symptoms (25). The child is 3 years old now and has not yet shown any symptoms. He is suggested to receive regular physical examination and possible treatment to maintain his plasma carnitine levels and prevent primary manifestations. Once the individual shows clinical manifestations of SPCD, he can get timely treatment. Here we verified one individual (ID: 84123) with reduced free carnitine deficiency but the value was normal at birth. The biochemical values of metabolites at birth may not reflect the real levels of the newborns themselves as the metabolites could be affected by the maternal concentration. In contrast, the results of genetic screening do not vary with age or diet. This case suggests that combining genetic screening with biochemical screening could reduce false-negative results.
According to previous studies in east of China, the prevalence of hyperphenylalaninemia was 1:11,572 and that of methylmalonic acidemia and homocystinuria (cblC type) was 1:28,000 (26,27). The carrier frequencies of mutations in PAH and MMACHC were consistent with the prevalence, indicating the rationality of our data. This is the first study to report the carrier frequencies of these genes in China.
In 2018, a pilot study of expanded carrier screening for 11 recessive diseases in China reported that the overall PAH carrier frequency of all ethnicities in China was 3.59%. The PAH frequency for Han ethnicity in that study was 3.30% (8), which was higher than that in our study. Our cohort were mainly born in Shanghai and the major population was the Han ethnicity in east of China. The Han population is distributed vastly in China, which may cause different carrier frequencies between the two studies. Another NBS study applied a molecular approach for four genetic diseases in Guizhou Province of southern China, including beta-thalassemia, G6PD deficiency, hyperphenylalaninemia, and non-syndromic hearing loss. The study selected 10 common mutations of PAH in the Chinese population, and reported a PAH carrier frequency of 0.78% among the 515 newborns (28) which was much lower than that in our study (1.77%). Our data revealed 20 newborns carrying 17 different mutations according to our criteria. We suggest that PAH is better screened as whole exons rather than as variants on several positions.
However, there is limitation for variants filtering in NBS as it is hard to assess the pathogenicity of some variants due to the inadequate information such as the absence of phenotype. Several diseases of high clinical importance are also technically challenging, making them difficult to assess with NGS. For example, Duchenne muscular dystrophy is typically caused by exonic rearrangements (29,30), whereas Silver-Russell syndrome is frequently caused by epimutations of H19/IGF2 imprinted domain (31,32). Some diseases such as spinal muscular atrophy have high homology (e.g., pseudogenes), special techniques are needed as pseudogenes are highly identical to the causative genes (33).
In conclusion, our study highlights the increased NBS efficiency of combining NGS screening with biochemical screening. This is the first study to investigate carrier frequencies of mutations in 77 genes causing IMDs in China.
Acknowledgments
We are very grateful to the newborns and their families as well as the clinicians participating in the study, and our laboratory team who contributed to this project.
Funding: This work was supported by the National Key R&D Program of China (2018YFC1002204 and 2019YFC1005100 to YY, 2018YFC1002400 to YS), the National Natural Science Foundation of China (81670812 and 81873671 to YY), the Jiaotong University Cross Biomedical Engineering (YG2017MS72 to YY), the Shanghai Municipal Commission of Health and Family Planning (201740192 to YY, 20184Y0073 to XL), the Shanghai Shen Kang Hospital Development Center new frontier technology joint project (SHDC12017109 to YY), the Shanghai Science and Technology Commission (19140904500 to YY), and the Shanghai Municipal Education Commission-Gaofeng Clinical Medicine Grant Support (20191908 to YY).
Footnote
Reporting Checklist: The authors have completed the STROBE reporting checklist. Available at http://dx.doi.org/10.21037/atm-20-1147
Data Sharing Statement: Available at http://dx.doi.org/10.21037/atm-20-1147
Peer Review File: Available at http://dx.doi.org/10.21037/atm-20-1147
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/atm-20-1147). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and was approved by Xinhua Hospital Ethics Committee Affiliated to Shanghai Jiao Tong University School of Medicine (XHEC-C-2017-021-2). Written consent was exempted as the residual DBS samples were reused in this study.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Ombrone D, Giocaliere E, Forni G, et al. Expanded newborn screening by mass spectrometry: New tests, future perspectives. Mass Spectrom Rev 2016;35:71-84. [Crossref] [PubMed]
- Holm IA, Agrawal PB, Ceyhan-Birsoy O, et al. The BabySeq project: implementing genomic sequencing in newborns. BMC Pediatr 2018;18:225. [Crossref] [PubMed]
- Berg JS, Agrawal PB, Bailey DB Jr, et al. Newborn Sequencing in Genomic Medicine and Public Health. Pediatrics 2017;139:e20162252. [Crossref] [PubMed]
- Ceyhan-Birsoy O, Murry JB, Machini K, et al. Interpretation of Genomic Sequencing Results in Healthy and Ill Newborns: Results from the BabySeq Project. Am J Hum Genet 2019;104:76-93. [Crossref] [PubMed]
- Meng L, Pammi M, Saronwala A, et al. Use of Exome Sequencing for Infants in Intensive Care Units: Ascertainment of Severe Single-Gene Disorders and Effect on Medical Management. JAMA Pediatr 2017;171:e173438. [Crossref] [PubMed]
- Petrikin JE, Cakici JA, Clark MM, et al. The NSIGHT1-randomized controlled trial: rapid whole-genome sequencing for accelerated etiologic diagnosis in critically ill infants. NPJ Genom Med 2018;3:6. [Crossref] [PubMed]
- Luo X, Wang R, Fan Y, et al. Next-generation sequencing as a second-tier diagnostic test for newborn screening. J Pediatr Endocrinol Metab 2018;31:927-31. [Crossref] [PubMed]
- Zhao S, Xiang J, Fan C, et al. Pilot study of expanded carrier screening for 11 recessive diseases in China: results from 10,476 ethnically diverse couples. Eur J Hum Genet 2019;27:254-62. [Crossref] [PubMed]
- Feuchtbaum L, Carter J, Dowray S, et al. Birth prevalence of disorders detectable through newborn screening by race/ethnicity. Genet Med 2012;14:937-45. [Crossref] [PubMed]
- Li W, Geng GX, Fan X, et al. Application of tandem mass spectrometry in screening of neonatal inherited metabolic diseases and result analysis. Maternal & Child Health Care of China 2015;30:5479-82.
- Wang LW, Ni M, Zhao DH, et al. Analysis of screening results of neonatal genetic metabolic diseases in Henan province by tandem mass spectrometry. Journal of Community Medicine 2019;17:1185-7.
- Yan HM, Jia ZJ, Liu J, et al. Tandem mass spectrometry screening of 565 182 newborns for inherited metabolic diseases in Hunan province. Chinese Journal of Applied Clinical Pediatrics 2019;34:1541-5.
- Zastrow DB, Baudet H, Shen W, et al. Unique aspects of sequence variant interpretation for inborn errors of metabolism (IEM): The ClinGen IEM Working Group and the Phenylalanine Hydroxylase Gene. Hum Mutat 2018;39:1569-80. [Crossref] [PubMed]
- Tang NL, Hwu WL, Chan RT, et al. A founder mutation (R254X) of SLC22A5 (OCTN2) in Chinese primary carnitine deficiency patients. Hum Mutat 2002;20:232. [Crossref] [PubMed]
- Boycott KM, Vanstone MR, Bulman DE, et al. Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat Rev Genet 2013;14:681-91. [Crossref] [PubMed]
- Cappellini MD, Fiorelli G. Glucose-6-phosphate dehydrogenase deficiency. Lancet 2008;371:64-74. [Crossref] [PubMed]
- Speiser PW, Arlt W, Auchus RJ, et al. Congenital Adrenal Hyperplasia Due to Steroid 21-Hydroxylase Deficiency: An Endocrine Society Clinical Practice Guideline. J Clin Endocrinol Metab 2018;103:4043-88. [Crossref] [PubMed]
- Jiang X, Tang F, Feng Y, et al. The adjustment of 17-hydroxyprogesterone cut-off values for congenital adrenal hyperplasia neonatal screening by GSP according to gestational age and age at sampling. J Pediatr Endocrinol Metab 2019;32:1253-8. [Crossref] [PubMed]
- Kopacek C, de Castro SM, Prado MJ, et al. Neonatal screening for congenital adrenal hyperplasia in Southern Brazil: a population based study with 108,409 infants. BMC Pediatrics 2017;17:22. [Crossref] [PubMed]
- Güran T, Tezel B, Gürbüz F, et al. Neonatal Screening for Congenital Adrenal Hyperplasia in Turkey: A Pilot Study with 38,935 Infants. J Clin Res Pediatr Endocrinol 2019;11:13-23. [Crossref] [PubMed]
- Srichomkwun P, Takamatsu J, Nickerson DA, et al. DUOX2 Gene Mutation Manifesting as Resistance to Thyrotropin Phenotype. Thyroid 2017;27:129-31. [Crossref] [PubMed]
- Wang F, Lu K, Yang Z, et al. Genotypes and phenotypes of congenital goitre and hypothyroidism caused by mutations in dual oxidase 2 genes. Clin Endocrinol (Oxf) 2014;81:452-7. [Crossref] [PubMed]
- Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405-24. [Crossref] [PubMed]
- Wang Y, Taroni F, Garavaglia B, et al. Functional analysis of mutations in the OCTN2 transporter causing primary carnitine deficiency: lack of genotype-phenotype correlation. Hum Mutat 2000;16:401-7. [Crossref] [PubMed]
- El-Hattab AW, Scaglia F. Disorders of carnitine biosynthesis and transport. Mol Genet Metab 2015;116:107-12. [Crossref] [PubMed]
- Gu X, Wang Z, Ye J, et al. Newborn screening in China: phenylketonuria, congenital hypothyroidism and expanded screening. Ann Acad Med Singap 2008;37:107-10. [PubMed]
- Wang F, Han L, Yang Y, et al. Clinical, biochemical, and molecular analysis of combined methylmalonic acidemia and hyperhomocysteinemia (cblC type) in China. J Inherit Metab Dis 2010;33 Suppl 3:S435-42. [Crossref] [PubMed]
- Huang S, Xu Y, Liu X, et al. Molecular newborn screening of four genetic diseases in Guizhou Province of South China. Gene 2016;591:119-22. [Crossref] [PubMed]
- Chen C, Ma H, Zhang F, et al. Screening of Duchenne muscular dystrophy (DMD) mutations and investigating its mutational mechanism in Chinese patients. PloS one 2014;9:e108038. [Crossref] [PubMed]
- Cho A, Seong MW, Lim BC, et al. Consecutive analysis of mutation spectrum in the dystrophin gene of 507 Korean boys with Duchenne/Becker muscular dystrophy in a single center. Muscle & nerve 2017;55:727-34. [Crossref] [PubMed]
- Ishida M. New developments in Silver-Russell syndrome and implications for clinical practice. Epigenomics 2016;8:563-80. [Crossref] [PubMed]
- Wu D, Gong C, Su C. Genome-wide analysis of differential DNA methylation in Silver-Russell syndrome. Science China Life sciences 2017;60:692-9. [Crossref] [PubMed]
- Kou H-S, Wang CC. Molecular inversion probes equipped with discontinuous rolling cycle amplification for targeting nucleotide variants: Determining SMN1 and SMN2 genes in diagnosis of spinal muscular atrophy. Analytica chimica acta 2017;977:65-73. [Crossref] [PubMed]


