
Tumor mutation burden in Chinese cancer patients and the underlying driving pathways of high tumor mutation burden across different cancer types
Introduction
High tumor mutation burden (TMB) has been associated with improved response to immune checkpoint inhibitors (ICIs) because elevated TMB increases the odds of generating immunogenic neoantigens (1,2). TMB was revealed to be an independent predictor of responses to ICIs not only in non-small cell lung cancer (NSCLC) (3), but also in small-cell lung cancer (SCLC) (4), melenoma (5), and other varieties of cancer (6). Multiple clinical trials have demonstrated the positive correlation between TMB and response to ICIs. KEYNOTE-001 has shown that in NSCLC patients receiving pembrolizumab, those with higher TMB had an improved overall response rate (ORR) and longer progression-free survival (PFS). TMB has been previously calculated by whole-exome sequencing (WES) (1,7). Nevertheless, its assessment by WES could be substantially limited by its high cost, the lack of deep coverage, and the additional bioinformatics demands. Multiple studies have reported that tatgeted sequencing panels containing coding regions of several hundreds of cancer-related genes can accurately estimate TMB and predict response to immunotherapy (8-11).
Although TMB may be a pan-cancer predictor for immune check point inhibitor, different tumors have different immune features and TMBs (12,13), and their potential driving mechanisms are different (14,15). For instance, deficiency in DNA damage response (DDR) pathway can raise the overall mutation burden in bladder cancer (16). In colon cancer, the mismatch repair (MMR)-deficient tumors were found to have a higher TMB than the MMR-proficient tumor (17). However, in breast cancers, tumors with mutations in BRCA1, a central gene in the homologous recombination pathway, exhibited a greater mutational burden than BRCA1-wt tumors (18). Given these diverse findings, further exploring the distinction of underlying driving pathway between different cancer types may be clinically significant.
Besides the disparity between cancer types, TMB may also vary across different ethnic populations. In NSCLC, for which targeted therapy was first engineered, a huge gap of efficacy was detected between Western populations and East Asian population in 2000s. This can be explained by the fact that East Asian populations harbor a higher percentage of epidermal growth factor receptor (EGFR) mutation (19-22). A similar ethnic diversity in relation to the EGFR mutation and NSCLC may also exist for TMB. However, most of the studies concerning TMB have been conducted in Western populations, and thus the TMB features in Chinese patients have not been well established. This may have great clinical significance for oncologists in the era of immune therapy, especially in China. Furthermore, if we can find a way to predict TMB with fewer combinations of genes, it will reduce the cost of sequencing and provide more convenience for clinicians.
In this study, we examined the TMB landscape of a cohort of 5,660 Chinese cancer patients, spanning 11 cancer types, using either a 295- or a 520-gene NGS panel. We established cancer-specific and histology-specific biological pathways associated with TMB status. In addition, as a proof of concept, an unsupervised algorithm was conducted using stepwise logistic regression to generate TMB-predicting signatures from both lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC).
Methods
Cohort selection and study design
We reviewed the genomic profiling data of 5,660 cancer patients from the following 9 participating centers: Changzheng Hospital, The Affiliated Hospital of Qingdao University, Fudan University Shanghai Cancer Center, Jiangsu Cancer Hospital & Jiangsu Institute of Cancer Research, The Affiliated Cancer Hospital of Nanjing Medical University, The First Affiliated Hospital of Suzhou University, The First Affiliated Hospital of Zhejiang University, Taizhou Central Hospital, Affiliated Hangzhou First People’s Hospital and Eastern Hepatobiliary Surgery Hospital. Samples were collected from April 2015-April 2018. There were 3 samples types: fresh tissue (n=2,177), formalin-fixed, paraffin-embedded (FFPE) (n=3,294) and pleural fluid (n=189), which were profiled in a Clinical Laboratory Improvement Amendments (CLIA)-certified sequencing laboratory (Burning Rock Biotech, Guangzhou, China) using the OncoScreen 295 (n=2,026) or OncoScreenPlus 520 (n=3,634) cancer-related gene panel. Of note, cases with maximal allelic frequency of less than 5% were not enrolled in this cohort. An external cohort consisting of 8,092 samples with WES sequencing data was downloaded from The Cancer Genome Atlas (TCGA) database to evaluate the in silico correlation of TMB using the 295- and 520-gene panels and WES. Eligible patients were histologically assessed according to the latest World Health Organization Criteria.
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and was approved by the Ethic Committee of Changzheng Hospital (2017SL016). Written informed consent was obtained from each patient for the use of their specimen.
NGS library preparation and sequencing
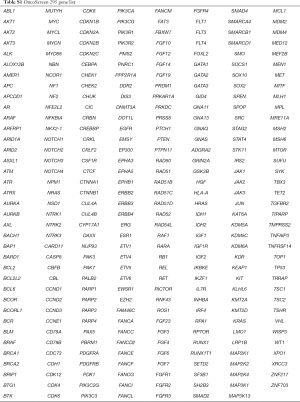
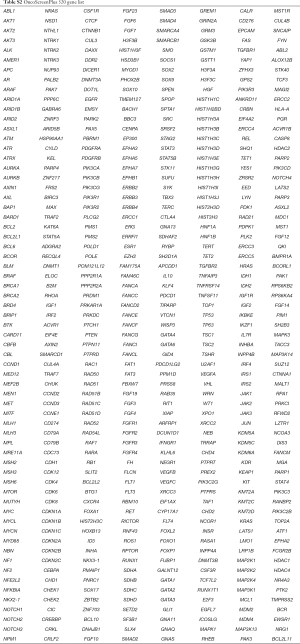
Capture-based targeted deep sequencing was performed using the 295- or 520-gene panel, spanning 1.44 and 1.64 Mb of the human genome, respectively. The gene list for each panel was listed in Tables S1 and S2. The detailed NGS library and sequencing protocol preparation was performed as previously described (23). In brief, DNA was fragmented by Covaris M220 focused ultrasonicator (Covaris, Inc., Woburn, MA, USA) followed by end repair, phosphorylation, dA addition, and adaptor ligation for library construction. Then, DNA library was purified by using Agencourt AMPure beads (Beckman Coulter, Fullerton, CA, USA). The quality and the size of the fragments were assessed using Qubit 2.0 fluorimeter with the dsDNA high-sensitivity assay kit (Life Technologies, Carlsbad, CA, USA). Indexed samples were sequenced on Nextseq500 (Illumina, Inc., USA) with paired-end reads.
Full table
Full table
TMB calculation and microsatellite instability (MSI) assessment
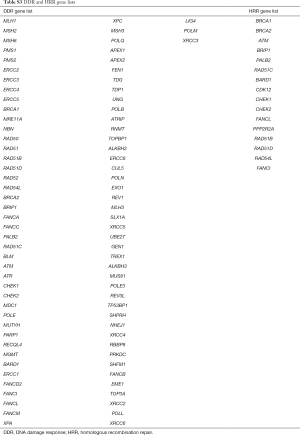
For sequencing data from the 295- or 520-gene panel, the somatic alterations in exons of coding regions and the adjacent 20-bp length of both upstream and downstream sequences were included in the calculation of TMB. The copy number variation and fusion were not counted. Alterations in the mutations of EGFR (exon 18–21) and ALK (amino acid 1,116–1,382) kinase domains were also excluded from the TMB calculation. A maximum allelic fraction (max.AF) of 5% was defined as the detection limit for TMB assessment using in-house validation, and samples with max.AF <5% were excluded. MSI status was determined as previously described (24). Additionally, homologous recombination repair (HRR) and DDR were defined as any non-synonymous mutation in the coding region of 16 and 87 genes, respectively. Detailed gene lists are provided in Table S3.
Full table
Analysis of the correlation of underlying pathways and TMB
To compute the significance of the correlation of each pathway with TMB, the patients were divided into two sub-groups: one group included those with any mutation in the specific pathway, and the other group included those without any such mutation. The ratio of the mean TMB of the patients with and without mutations in this pathway was calculated as the main statistical indicator. Next, regions with the same size covered by all genes from each pathway were randomly selected from our panel with 1,000 repetitions to simulate the distribution of the statistic and compute the significance, while controlling for bias in which a high-TMB sample could elevate the number of mutations among any set of genes. In each simulation, patients were also divided into two sub-groups mutated or non-mutated, based on the mutation status of the randomly selected regions, and the mean TMB ratios of these two groups were also calculated.
Gene signature development for TMB prediction
A machine learning algorithm was used in the cohorts with LUAD and LUSC to construct TMB prediction models. Samples of 300 patients with LUAD and 100 patients with LUSC were selected randomly from the entire cohort as independent test sets. The remaining samples, utilized as training sets, were used to establish the TMB class prediction model. To select the most predictive genes, a t-test was employed firstly in the training set to find the genes related to TMB as candidate genes. Then, the CfsSubsetEval attribute evaluator and the BestFirst search method of WEKA software (version 3.8) were used for feature selection (25). The predictive capability of each attribute and the degree of redundancy between two different attributes were measured using the CfsSubsetEval attribute evaluator. Furthermore, a set of attributes with a high correlation and low-coupling was generated. The BestFirst search method searched the feature subset space through a greedy hill-climbing strategy augmented with a backtracking facility. Next, to avoid over-fitting, a ten-fold cross-validation was utilized in the feature selection procedure. Considering the convenience of clinical application, logistic regression was used to establish the TMB class prediction model by gene features. To evaluate the performance of the model, both ten-fold cross-validation of the training dataset and independent test datasets were utilized.
Statistical analysis
All data, except for the feature selection step of machine learning, were analyzed using Software R (Version 3.4.0). The correlation between TMB (as calculated by the 295- and 520-gene panels) and WES was evaluated by linear regression. Wilcoxon signed-rank test was used to compare the mutation loads between the age groups among the TMB-high, -medium, and -low patients. Comparisons between the mutation burden in male and female patients, MSI-H and microsatellite-stable (MSS) patients, DDR deficient and DDR proficient patients, and HRR deficient and HRR proficient patients were also performed using the Wilcoxon signed-rank test. For all statistical tests, a P value <0.05 was considered statistically significant.
Results
The landscape of TMB across different cancer types in the Chinese population
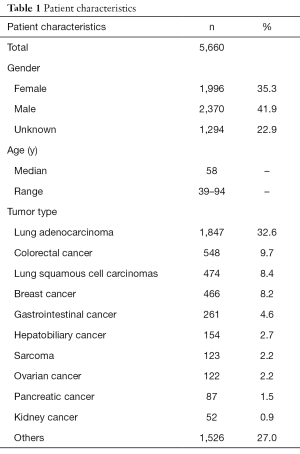
This cohort contained 1,996 (35.3%) females and 2,370 (41.9%) males, and gender information of 1,294 (22.9%) cases was unavailable. Median age of these patients was 58 years, ranging from 39 to 94 years (Table 1). For subsequent analyses, patients of this cohort with 11 distinct cancer and histology types, were classified into the following 3 main types on the basis of tumor origin and evolution: LUAD (1,847/5,660, 32.6%), colorectal cancer (548/5,660, 9.7%), and LUSCs (474/5,660, 8.4%). Other cancer types included breast cancer (466/5,660, 8.2%), gastrointestinal cancer (261/5,660, 4.6%), hepatobiliary cancer (154/5,660, 2.7%), etc. The last cancer type group, “others” (n=1,526/5,660, 27.0%), included cancer types containing less than 50 unique specimens (n=417), lung cancers except for LUAD and LUSC (n=899), and cases with unknown cancer types (n=210).
Full table
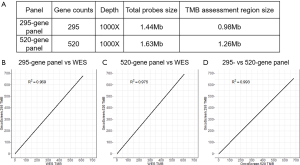
Detailed panel information is presented in Figure S1A. The TMB, assessed by the 295- and 520-gene panels and WES closely correlated with each other (295-gene panel vs. WES, R2 =0.969; 520-gene panel vs. WES, R2 =0.975; 295- vs. 520-gene panel, R2 =0.993; Figure S1B,C,D). These results indicated that the comprehensive genomic profiling using 295- and 520-gene panels can accurately reveal the actual mutation burden.
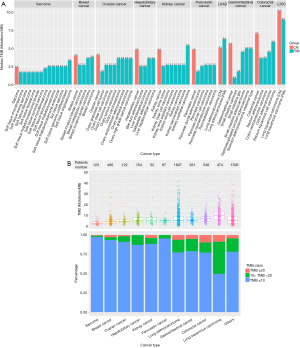
We performed comparative mutation burden analysis between our Chinese study cohort and a larger cohort (over 100,000 samples) reported by Foundation Medicine Inc. (FMI) (8). In our cohort, the TMB distribution was highly variable between and within cancer classes, ranging from 0 to 723.8 mutations/Mb, with a median TMB of 5.6 mutations/Mb. The median TMB was slightly higher than that from the FMI dataset, which was 3.6 mutations/Mb. Overall, 5.4% (n=305) of the patients had a TMB higher than 20 mutations/Mb, 16.5% (n=936) cases had a TMB between 10 and 20 mutations/Mb, and 78.1% (n=4,439) cases had a TMB of less than 10 mutations/Mb in our cohort.
Among all the cancer groups, sarcomas had the lowest mutation burden (median TMB 2.4 mutations/Mb) in our cohort, which agreed well with the FMI results (median of each sarcoma subtypes ranged from 1.7 to 3.3 mutations/Mb). The median TMB of breast cancer ranked second in our cancer groups in terms of TMB from low to high and coincided with that of the FMI population (median of each breast cancer subtypes ranged from 2.7 to 3.8 mutations/Mb). As to ovarian cancer, the median TMB in our cohort was 4.1 mutation/Mb, and the range of median TMB for each ovarian cancer subtype in the FMI dataset was 1.8–3.6 mutation/Mb. We found that the median TMB of hepatobiliary cancer, kidney cancer, and pancreatic cancer was the same in our cohort (4.8 mutations/Mb), and higher than that in the FMI dataset (hepatobiliary cancer median =2.5–3.6 mutations/Mb; kidney cancer median = from 2.5–5.4 mutations/Mb; pancreatic cancer median =1.8–2.7 mutations/Mb). In gastrointestinal cancer and colorectal cancer, the median TMB was 5.6 and 7.1 mutations/Mb, respectively, higher than those of the FMI population (gastrointestinal cancer median =0.9–5.0 mutations/Mb; colorectal cancer median 3.6–5.9 mutations/Mb). In addition, cancers related to chronic mutagen exposures such as lung cancers exhibited greater hyper-mutation than other cancer groups in our cohort. Within lung cancers, LUSC was more highly mutated than LUAD (median 10.2 vs. 5.1 mutations/Mb), and consistent with conclusions from the FMI population (median 9.0 vs. 6.3 mutations/Mb) (Figure 1A,B).
Association between TMB and demographic/molecular features
TMB-medium (63 years) and high groups (63 years) were significantly older than the TMB-low group (56 years; P<0.001, Wilcoxon signed-rank test; n=4,328; Figure 2A). This phenomenon was also observed in the lung cancer subpopulation (high TMB, median age =65 years; medium TMB, 63 years; low TMB, 59 years; P<0.001, n=1,801; Figure 2B), which was the major tumor type in this study. Furthermore, our analysis revealed that male patients more commonly correlated with higher TMB than the female patients, with statistical significance (median TMB 6.3 vs. 4.0 mutations/Mb, P<0.001, n=4,366; Figure 2C), in both the whole cohort and the lung cancers group (median TMB 7.1 vs. 4.0 mutations/Mb, P<0.001, n=1,810; Figure 2D).
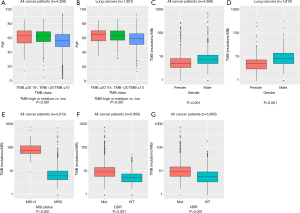
We further established that the MSI-high patients usually had a higher TMB than the MSS patients (median TMB 71.4 vs. 5.1 mutations/Mb, P<0.001, n=4,513, Figure 2E). Alterations in DDR occurred in all 11 cancer type groups, with alteration frequencies ranging from 26.4% (23/87) in pancreatic cancer to 57.8% (274/474) in LUSC. We observed that DDR-deficient patients had a significantly higher TMB than the DDR-proficient patients (median TMB 7.9 vs. 4.1 mutations/Mb, P<0.001, n=5,660, Figure 2F).
Similar to DDR, HRR alterations were identified in the patients of all 11 cancer type groups, with a minimal alteration frequency of 11.5% (6/52) in kidney cancer and a maximal frequency of 34.5% (161/466) in breast cancer. HRR-deficient patients had a significantly higher TMB than HRR-proficient patients (median TMB 8.2 vs. 4.8 mutations/Mb, P<0.001, n=5,660, Figure 2G).
Underlying driving pathways of high TMB across different cancer types
We investigated the distribution of mutations across Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways in different cancer groups (Figure 3A). The minimal percentage of mutated cases was observed in the MMR pathway of pancreatic cancer (0/87, 0%), whereas the maximal percentage occurred in the PI3K-Akt signaling pathway of colorectal cancer (537/548, 98.0%).
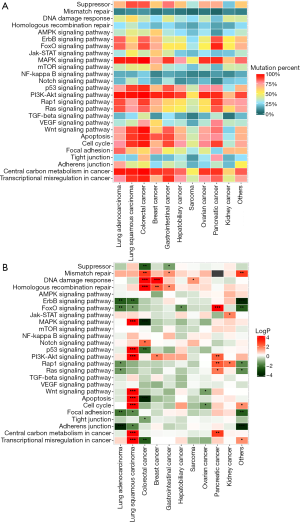
Some of the pathways displayed significant association with TMB status in different cancer groups, but no pathway had a universal association with TMB. Moreover, we observed that an alteration in an identical pathway but in different cancer groups may indicate an opposite direction of the TMB status (Figure 3B, Figure S2).

TMB predictive signature (TPS) development
Molecular signatures consisting of 23 and 16 gene features were derived for TMB status prediction in LUAD and LUSC, respectively (Figure 4A). In LUAD, 22 gene features were positively correlated with the TMB status, with a correlation coefficient value ranging from 0.34 for ATR to 1.63 for LRP1B. Only EGFR (oncogenic driver variants) was negatively correlated to TMB (correlation coefficient =−1.13). In LUSC, all 16 identified gene features were positively associated with TMB. Among them, KMT2A was the most highly correlated with the TMB status, with a correlation coefficient of 1.67, followed by TP53 (1.60) and RUNX1T1 (1.59).
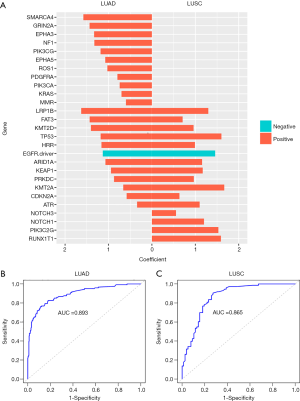
The TMB predicted by TPS was in remarkable agreement with the TMB directly calculated by the NGS panels, as measured by area under the curve (AUC) (LUAD, AUC =89.3%, Figure 4B; LUSC, AUC =86.5%, Figure 4C) and seven other parameters (Table S4) in both NSCLC subtypes.

Full table
Discussion
We characterized the landscape of TMB in a cohort of 5,660 Chinese cancer patients across 11 cancer groups. To our knowledge, our cohort is the largest reported Chinese cohort concerning TMB in a pan-cancer population. We observed a rich variation in mutational burden across and within cancer types, which was consistent with previous studies (1,7,8). Patients with high TMB can be identified in nearly all cancer types, implying that patients with any cancer types may have potentially benefit from immunotherapy. In our study cohort, the median TMB of several tumor types was higher than that of the FMI dataset. Several factors can account for these results including but not limited to the difference in ethnicity, age, stage, line of treatment, cohort size, and TMB calculation algorithm.
Numerous previous studies have explored the demographic and molecular features associated with TMB and yielded conflicting findings. A recent study in a Chinese population reported the absence of a correlation between TMB and age or gender, but only included 16 adolescent patients (9). However, an investigation in a Caucasian population established that a high TMB was related to older age, but no difference in the median TMB existed between female and male patients (8). In our cohort, the higher TMB was correlated with older age and male gender. We also revealed that MSI-high, DDR, and HRR deficiency commonly indicated a higher TMB than MSS, DDR proficiency, and HRR proficiency, which is consistent with the findings of previous studies (8).
Increasing evidence suggests that the underlying TMB-associated biological mechanisms vary across different cancer types. DDR deficiency leads to a high TMB in bladder cancer, whereas MMR deficiency leads to hypermutation in colon cancer (16,17). Here, we estimated the association of 26 crucial biological pathways and the TMB status in 11 tumor type groups. Besides the pathways related to genomic instability and DNA repair, such as MMR, HRR, and DDR, signaling pathways were also included in our analysis. The correlation between TMB and the biological pathways was found to be both cancer- and histology-specific. LUSC is characterized by a high mutation burden and marked genomic complexity (26). There are frequent alterations of CDKN2A, RB1, and AKT in LUSC, which are involved in the following pathways: cell cycle control, p53 signaling pathway, apoptosis, PI3K-Akt pathway, central carbon metabolism, and MAPK signaling pathway (27). The frequent alterations in these pathways in LUSC are the potential underlying biological basis for a high TMB, which is consistent with our results that all the above-mentioned pathways are correlated with a high TMB in LUSC.
Notch signaling pathway was correlated with a low-TMB status in pancreatic cancer. This finding is in agreement with those of previous studies reporting that aberrant Notch signaling was involved in tumor initiation and tumor maintenance in pancreatic cancer (28,29), and patients with pancreatic cancer commonly had a low TMB (7,26). Nevertheless, it is worth noting that we have not definitively demonstrated the causality between mutated pathways and the mutation burden.
Efforts have been previously made to identify gene alterations associated with an increased TMB (10). Herein, we generated 23- and 16-gene signatures in LUAD and LUSC, respectively, to establish the TMB, reaching an accuracy of 88.4% (LUAD) and 79.3% (LUSC), respectively. To date, these are the smallest gene sets reported for TMB prediction.
Conclusions
This study is the largest pan-cancer NGS sequencing cohort reported in a Chinese population to date. In this study, using the 295- and 520-gene NGS panels, we produced a TMB estimation which strongly correlated with those calculated by WES. Using our targeted sequencing panel, we revealed the diversity of TMB between the Chinese and Caucasian populations, identified drivers and predictors of TMB status, and found highly diverse patterns across different cancer types. Moreover, gene signatures consisting of 23 and 16 genes were derived for TMB status prediction in LUAD and LUSC, respectively, with only 12 genes shared by both subtypes, suggesting that the two NSCLC histological subtypes possess distinct underlying mechanisms for induction of the TMB status.
Our findings extend the knowledge of the diversity across different ethnicities and reveal the underlying biological mechanisms for high TMB. These results might be clinically significant, especially for physicians in China, and may be helpful for developing more precise and accessible TMB assessment panels and algorithms in more cancer types.
Acknowledgments
Funding: None.
Footnote
Data Sharing Statement: Available at http://dx.doi.org/10.21037/atm-20-3807
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/atm-20-3807). Dr. SC, Dr. YSZ, Dr. HHZ, Dr. JY and Dr. TH report that they are employees of Burning Rock Biotech. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and was approved by the Ethic Committee of Changzheng Hospital (2017SL016). Written informed consent was obtained from each patient for the use of their specimen.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Alexandrov LB, Nik-Zainal S, Wedge DC, et al. Signatures of mutational processes in human cancer. Nature 2013;500:415-21. [Crossref] [PubMed]
- Addeo A, Weiss GJ. Measuring tumor mutation burden in cell-free DNA: advantages and limits. Transl Lung Cancer Res 2019;8:553-5. [Crossref] [PubMed]
- Hellmann MD, Ciuleanu TE, Pluzanski A, et al. Nivolumab plus Ipilimumab in Lung Cancer with a High Tumor Mutational Burden. N Engl J Med 2018;378:2093-104. [Crossref] [PubMed]
- Hellmann MD, Callahan MK, Awad MM, et al. Tumor Mutational Burden and Efficacy of Nivolumab Monotherapy and in Combination with Ipilimumab in Small-Cell Lung Cancer. Cancer Cell 2018;33:853-61.e4. [Crossref] [PubMed]
- Eroglu Z, Zaretsky JM, Hu-Lieskovan S, et al. High response rate to PD-1 blockade in desmoplastic melanomas. Nature 2018;553:347-50. [Crossref] [PubMed]
- Goodman AM, Kato S, Bazhenova L, et al. Tumor Mutational Burden as an Independent Predictor of Response to Immunotherapy in Diverse Cancers. Mol Cancer Ther 2017;16:2598-608. [Crossref] [PubMed]
- Lawrence MS, Stojanov P, Polak P, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 2013;499:214-8. [Crossref] [PubMed]
- Chalmers ZR, Connelly CF, Fabrizio D, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med 2017;9:34. [Crossref] [PubMed]
- Zhuang W, Ma J, Chen X, et al. The Tumor Mutational Burden of Chinese Advanced Cancer Patients Estimated by a 381-cancer-gene Panel. J Cancer 2018;9:2302-7. [Crossref] [PubMed]
- Roszik J, Haydu LE, Hess KR, et al. Novel algorithmic approach predicts tumor mutation load and correlates with immunotherapy clinical outcomes using a defined gene mutation set. BMC Med 2016;14:168. [Crossref] [PubMed]
- Rizvi H, Sanchez-Vega F, La K, et al. Molecular Determinants of Response to Anti-Programmed Cell Death (PD)-1 and Anti-Programmed Death-Ligand 1 (PD-L1) Blockade in Patients With Non-Small-Cell Lung Cancer Profiled With Targeted Next-Generation Sequencing. J Clin Oncol 2018;36:633-41. [Crossref] [PubMed]
- Abida W, Cheng ML, Armenia J, et al. Analysis of the Prevalence of Microsatellite Instability in Prostate Cancer and Response to Immune Checkpoint Blockade. JAMA Oncol 2019;5:471-8. [Crossref] [PubMed]
- Wu YM, Cieslik M, Lonigro RJ, et al. Inactivation of CDK12 Delineates a Distinct Immunogenic Class of Advanced Prostate Cancer. Cell 2018;173:1770-82.e14. [Crossref] [PubMed]
- Pan F, Wingo TS, Zhao Z, et al. Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells. Nat Commun 2017;8:15102. [Crossref] [PubMed]
- Humphris JL, Patch AM, Nones K, et al. Hypermutation In Pancreatic Cancer. Gastroenterology 2017;152:68-74.e2. [Crossref] [PubMed]
- Yap KL, Kiyotani K, Tamura K, et al. Whole-exome sequencing of muscle-invasive bladder cancer identifies recurrent mutations of UNC5C and prognostic importance of DNA repair gene mutations on survival. Clin Cancer Res 2014;20:6605-17. [Crossref] [PubMed]
- Stadler ZK, Battaglin F, Middha S, et al. Reliable Detection of Mismatch Repair Deficiency in Colorectal Cancers Using Mutational Load in Next-Generation Sequencing Panels. J Clin Oncol 2016;34:2141-7. [Crossref] [PubMed]
- Nolan E, Savas P, Policheni AN, et al. Combined immune checkpoint blockade as a therapeutic strategy for BRCA1-mutated breast cancer. Sci Transl Med 2017;9:eaal4922.
- Calvo E, Baselga J. Ethnic differences in response to epidermal growth factor receptor tyrosine kinase inhibitors. J Clin Oncol 2006;24:2158-63. [Crossref] [PubMed]
- Blackhall F, Ranson M, Thatcher N. Where next for gefitinib in patients with lung cancer? Lancet Oncol 2006;7:499-507. [Crossref] [PubMed]
- Shigematsu H, Lin L, Takahashi T, et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J Natl Cancer Inst 2005;97:339-46. [Crossref] [PubMed]
- Sun Y, Ren Y, Fang Z, et al. Lung adenocarcinoma from East Asian never-smokers is a disease largely defined by targetable oncogenic mutant kinases. J Clin Oncol 2010;28:4616-20. [Crossref] [PubMed]
- Mao X, Zhang Z, Zheng X, et al. Capture-Based Targeted Ultradeep Sequencing in Paired Tissue and Plasma Samples Demonstrates Differential Subclonal ctDNA-Releasing Capability in Advanced Lung Cancer. J Thorac Oncol 2017;12:663-72. [Crossref] [PubMed]
- Zhu L, Huang Y, Fang X, et al. A Novel and Reliable Method to Detect Microsatellite Instability in Colorectal Cancer by Next-Generation Sequencing. J Mol Diagn 2018;20:225-31. [Crossref] [PubMed]
- Frank E, Hall M, Trigg L, et al. Data mining in bioinformatics using Weka. Bioinformatics 2004;20:2479-81. [Crossref] [PubMed]
- Lee CH, Yelensky R, Jooss K, et al. Update on Tumor Neoantigens and Their Utility: Why It Is Good to Be Different. Trends Immunol 2018;39:536-48. [Crossref] [PubMed]
- Cancer Genome Atlas Research N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 2012;489:519-25. [Crossref] [PubMed]
- Abel EV, Kim EJ, Wu J, et al. The Notch pathway is important in maintaining the cancer stem cell population in pancreatic cancer. PLoS One 2014;9:e91983. [Crossref] [PubMed]
- Garrido-Laguna I, Hidalgo M. Pancreatic cancer: from state-of-the-art treatments to promising novel therapies. Nat Rev Clin Oncol 2015;12:319-34. [Crossref] [PubMed]


