
Downregulated placental expression of linc00468 contributes to trophoblast dysfunction by inducing epithelial-mesenchymal transition
Introduction
PE is the most common idiopathic disorder with high prevalence, leading to approximately 60,000 worldwide pregnancy-related deaths worldwide each year (1). Because of the specific pathological changes in PE, it contributes to small systemic vasospasm, endothelial injury and ischemia, further causing adverse organ failure. Meanwhile, unfavorable pregnancy outcomes may occur, including postpartum hemorrhage and disseminated intravascular coagulation (DIC), premature birth, fetal growth restriction and placental abruption (2,3). To date, there is a lack of effective clinical treatments are lacking and delivery is the only solution to halt progression (4).
PE mainly originates from the placenta. Pathological studies have reported that the pathogenesis of PE is due to the failure of uterine spiral artery remodeling and is closely related to the function of trophoblasts. Aberrant extravillous trophoblast (EVTs) phenotypes, such as abnormal trophoblast apoptosis (5), impaired EVT proliferation (6), and reduced EVT migration and invasion ability (7), which can increase blood flow resistance and cause hypoperfusion in this process, further impair the balance of the maternal-fetal interface (8). The trophoblastic cell columns are formed by this physiological conversion, which is called the epithelial-mesenchymal transition (EMT) (9). In placental development, patients with PE are characterized by the high expression of the epithelium, like E-cadherin and β-catenin and low expression of the mesenchyme, like N-cadherin, Vimentin, Snail, which facilitate the progression of extensive proliferation, migration and invasion into the maternal decidua and vessels (9,10).
Based on the advancements in genome-wide analysis of gene expression, non-coding RNA, especially long non-coding RNAs (lncRNAs), have emerged as new emerging players demonstrating latent roles in various human diseases (11,12), including PE (13-15). For example, the dysregulation of the lncRNA HIF1A antisense RNA 2 may participate in the function of trophoblast cell invasion and cell-cycle progression through binding to LSD1 to downregulate PHLDA1 expression (16). Recently, growing number of studies have revealed that lncRNAs regulate the EMT in PE, although the mechanism is still unknown. In our previous study, we investigated how resveratrol (RE) can affect the invasive ability of maternal trophoblasts by activating the EMT progress and may induce the Wnt/β-catenin signaling pathway in a PE mouse model (17). Therefore, in-depth exploration of the molecular mechanisms of lncRNAs may hold the potential for clinical strategies for treating PE (18). However, it is not yet fully understood how a dysregulated EMT causes PE.
In this study, we uncovered a novel linc00468 (19), which is markedly downregulated in most PE placental samples compared to normal placental samples. Our results showed that linc00468 could promote trophoblast phenotypic changes by inducing the EMT in EVTs. These results impelled us to identify the underlying signaling pathways that mediate this effect in PE.
Methods
Tissue collection and ethics statement
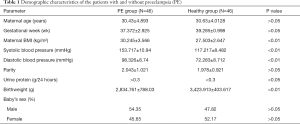
We obtained PE placental and normotensive placental tissues from 46 patients who underwent cesarean deliveries at the Jiangsu Women and Children Health Hospital between 2018 and 2019 (Nanjing, Jiangsu, China). We initially used sterile phosphate-buffered saline to wash all collected placental samples to remove any blood, and then quickly the samples stored in liquid nitrogen until we could extract protein and RNA. All procedures performed in this research were conducted in compliance with the principles of the Declaration of Helsinki. Risks and changes in the clinicopathological characteristics of all PE patients are recorded and presented in Table 1. The use of human placental samples was approved with written informed consent from the patients and authorized by the Ethics Committee of The First Affiliated Hospital of Nanjing Medical University.
Full table
Cell culture and transfection assay
HTR-8/SVneo, the human first-trimester EVT cell line, was generously provided by Dr. Charles Graham, Queen’s University, Canada and was cultured in DMEM (KEYGEN, Nanjing, China) medium supplemented with 10% fetal bovine serum, and 100 U/mL penicillin. The JAR, Jeg3, and BeWo cell lines were purchased from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China) and cultured according to the manufacturer’s instruction. JAR cells were cultured in DMEM (KEYGEN, Nanjing, China). JEG3 cells were also cultured in DMEM (KEYGEN, Nanjing, China) and BeWo cells were cultured in DMEM/F12K medium. All cell lines were grown under standard humidified air conditions at 37 °C with 5% CO2.
Cell transfection
Three individual small interfering RNAs (siRNA)- si-linc00468-#1, si-linc00468-#2 and si-linc00468-#3, and a negative control siRNA (si-NC) were acquired from Invitrogen. The SiRNA sequences were si-lin c00468 #1 (5'-UCAAGGACUUCAUUGGUGCAGCUGA-3'), si-linc00468 #2 (5'-AAAUCACGAACACCGAAAGAAGCCA-3'), si-linc00468 #3 (5'-UCAGCUGCACCAAUGAAGUCCUUGA-3'). Full-length cDNA of linc00468 (903 bp, NR_037593.1) was synthesized and cloned into the pcDNA3.1(+) mammalian expression plasmid vector (Invitrogen, USA). When the trophoblast cells reached 50% confluence, transfection was conducted with Lipofectamine 3000 transfection reagents (Life Technologies, Invitrogen, USA) according to the manufacturer’s instructions. Cells were harvested for subsequent assays at 48 hours post-transfection.
Quantitative real-time PCR (qRT-PCR)
We used Trizol reagent to isolate total RNA from cells. The RNA samples were reliably amplified and labeled to generate cDNA with Primer Script RT Master Mix (Takara, China) based on the provided directions in ABI prism 7500 system. PowerUp™ SYBR™ Green Master Mix (Applied Biosystems™, Invitrogen) was used for qRT-PCR following the manufacture’s protocol. GAPDH served as loading control and fold changes of each gene were calculated using the 2−ΔΔCt method (cycle threshold). The primers used were as follows: GAPDH (F:GAAGGTGAAGGTCGGAGTC, R:GAAGATGGTGATGGGATTTC); Linc00468 (F:CAAATCGTAGCCAGACCCGA, R:GCGAAGGAAGCATTTCGAGC); E-cadherin (F:TCCCATCAGCTGCCCAGAAA, R:TGACTCCTGTGTTCCTGTTA); β-catenin (F:CTCAGGACAAGGAAGCTGCAGAAGC, R:CAAGGCATCCTGGCCATATCCA) N-cadherin (F:CCCTGCTTCAGGCGTCTGTA, R:TGCTTGCATAATGCGATTTCACC); Vimentin (F:TCTACGAGGAGGAGATGCGG, R:GGTCAAGACGTGCCAGAGAC); Snail (F:GACCACTATGCCGCGCTCTT, R:TCGCTGTAGTTAGGC TTCCGATT). Each sample was tested in triplicate.
Cell proliferation assays
A cell counting kit-8 (CCK-8, MCE, USA) was used to assess cell viability. Ten microliters of the CCK-8 solution were added to each well of the plate after incubation for 0, 24, 48, 72, or 96 h. The data were obtained using a microplate reader and plotted against time to achieve growth curves. For colony formation experiments, cells were cultured for about 1 week until macroscopic colonies appeared. The colonies were then fixed with methanol and stained with crystal violet (Sigma, China) for 30 min separately.
For the 5-ethynyl-2'-deoxyuridine (EdU) experiment, cells were incubated with 50 mM Edu labelling medium (Ribobio, Guangzhou, China) for 2 h at 37 °C, 5% CO2 and then fixed in 4% paraformaldehyde for 30 min at room temperature. Then, cells were permeabilized with 0.5% Triton X-100. Finally, we used DAPI to localize the nucleus of cells. Proliferating cells were detected with a fluorescence microscope.
The Transwell migration and invasion assay
The cell invasion and migration assays were conducted using a Transwell chamber (8-µm pore size, Millipore, Billerica, MA, USA). Briefly, 5×104 cells were seeded in the upper chamber in serum-free medium and RPMI1640 medium containing 10% FBS was added to the lower chamber. After incubating for 24–48 h, the migrated or invading cells were fixed with paraformaldehyde 0.1% crystal violet. Next, cells that penetrated the membrane were observed under a light microscope and the number of cells from five fields was analyzed at random. All experiments were repeated in triplicate.
Flow cytometric analysis
Cells were cultured for 24 h at a density of 3×105 cells in a 6-well plate. HTR-8/SVneo and JAR cells transfected with si-linc00468 #2 or #3 or pcDNA3.1-linc00468 were harvested 48 h post-transfection by trypsinization. Then, double staining with fluorescein isothiocyanate (FITC)-Annexin V and propidium iodide (PI) was performed using the FITC Annexin V Apoptosis Detection Kit (BD Biosciences, Franklin Lakes, NJ, USA) according to the manufacturer’s recommendations. The results were analyzed via flow cytometry (FACScan; BD Biosciences) to determine the extent of apoptosis. For the cell-cycle experiment, cells were stained with PI (KGA214, Nanjing, China), and detected using a FACScan flow cytometer (BD Biosciences) to categorize and compare the percentages of cells in the G0–G1, S, and G2–M phases, followed by flow cytometric analysis. The tests were repeated in three independent experiments and are presented as the mean ± standard deviation (SD).
Western blot analysis
Total protein was lysed in pre-chilled RIPA protein extraction reagent and quantified using a bicinchoninic acid (BCA) protein assay kit (Beyotime Biotechnology, Shanghai, China). Protein was separated by SDS-PAGE and then transferred to 0.22-µm polyvinylidene difluoride PVDF membranes (Millipore, USA). Subsequently, the PVDF membranes were blocked with 5% non-fat milk in TBST buffer for 1.5 h at room temperature and then incubated with primary antibodies (E-cadherin, N-cadherin, Vimentin, Proteintech) and anti-GAPDH antibody (Santa Cruz, CA, USA) overnight at 4 °C. After washing, the secondary antibody, horseradish peroxidase-conjugated goat anti-mouse or goat anti-rabbit IgG (Proteintech) was added and incubation took place for 2 h. Enhanced chemiluminescence (Thermo) was used to detect densitometry by using the Quantity One software (Bio-Rad). GAPDH was used as an internal control. All of the experiments were conducted in triplicate.
Fluorescence in situ hybridization (FISH)
FISH was performed to detect the location of lincRNA. HTR-8/SVneo cells (6×104) were seeded in a 24-well plate and fixed in 4% paraformaldehyde for 30 min. Probe detection and DNA staining were conducted using the RiboTM Fluorescence in Situ Hybridization Kit, according to the manufacturer’s instructions. A red fluorescent probe was used to identify linc00468 and 4, 6-diamidino-2-phenyl-indole (DAPI) was used to label nuclei. The sequence of the RNA-FISH probe for linc00468 is 5’-AGCCCACCAGCCCTCCGCACCGTGGGCGAC-3’.
Immunofluorescence staining
HTR-8/SVneo cells were fixed in 4% paraformaldehyde for 5 min and permeated in 0.3% Triton X for 5 min. After washing by PBS, cells were incubated with 2% BSA at room temperature for 1 h. Following this, cells were incubated overnight with primary antibodies (E-cadherin, 1:50 dilution, Mouse mAB, 4A2, CST; N-cadherin, 1:200 dilution, Rabbit mAb, D4R1H, CST; Vimentin, 1:500 dilution, Rabbit mAb, D21H3, CST). Following the addition of secondary antibody, cells were incubated with DAPI for 5 min. An inverted fluorescence microscope was used to record images.
Statistical analysis
Data were obtained from three independent experiments. GraphPad Prism (version 7.0, La Jolla, CA, USA) software was used for statistical analysis. All measurement data are expressed as mean ± SD. Significant differences between two groups were analyzed using a paired t-test and were defined as follows: *, P<0.05; **, P<0.01; and ***, P<0.001.
Results
Expression of Linc00468 is downregulated in PE placentas
The expression levels of linc00468 were detected in 46 pairs of PE and corresponding normal placental samples by qRT-PCR. The results revealed that linc00468 expression levels were markedly downregulated in PE tissues (Figure 1A,B). We also summarized the patients’ demographic characteristics in Table 1. There were significant differences in maternal BMI, systolic blood pressure, diastolic blood pressure, urine protein and birthweight between the PE and the healthy samples taking statistical analysis into consideration (P<0.01, P<0.01, P<0.01, P<0.05 and P<0.01, respectively). On the contrary, there was no statistical difference between PE placentas and normal placentas in maternal age, gestational week, parity and baby’s sex (P>0.05). In addition, we discovered that the expression of linc00468 was 1.72-fold lower in the PE group compared with the healthy group (Figure 1B).
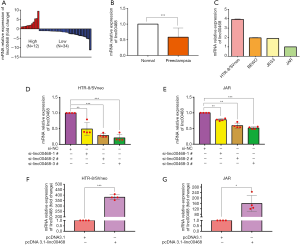
Knockdown and overexpression of linc00468 in trophoblast cell lines
To gain a deeper insight into the molecular function of linc00468, we first detected the expression of linc00468 in several trophoblast cell lines, including HTR-8/SVneo, BeWo, JEG-3 and JAR. As shown in Figure 1C, we found that the relative level of linc00468 in HTR-8/Svneo and JAR cell lines was respectively higher and lower than those in the other two cell lines. Thus, these two cell lines were chosen for subsequent exploration. Next, we synthesized three different siRNAs to knock down linc00468 in trophoblast cell lines. Forty-eight hours after the transfection, qRT-PCR results indicated that the use of si-linc00468 #2 and #3 was more efficient in silencing the expression of linc00468 in these two cell lines than si-linc00468 #1 (Figure 1D,E). Furthermore, linc00468 expression was upregulated by transfecting HTR-8/Svneo and JAR cell lines with the pcDNA 3.1-linc00468 expression vector (Figure 1F,G).
The effects of linc00468 on the proliferation and apoptosis of trophoblast cells
The sufficient silencing of linc00468 expression attenuated cell viability in HTR-8/Svneo and JAR cells, as measured by the cell-counting kit-8 (CCK8), which provides a colorimetric format to monitor trophoblast cell proliferation (Figure 2A,B). Next, we conducted flow cytometric trials to activate caspase proteases and cause mitochondrial dysfunction, which further indicated that overexpression of linc00468 ameliorated the apoptosis rate (Figure 2C,D,E). Results of microscopic ethynyl deoxyuridine (EdU) staing (Figure 2F,G) revealed that linc00468 exerted a suppressive effect to accelerate cell proliferation. As shown in Figure S1, cell viability was inhibited by si-linc00468 #2 and #3 transfection in the colony formation assay (Figure S1). These data taken together illustrate that linc00468 may exert an effect on cell proliferation and apoptosis, indicating that knockdown of linc00468 plays a key role in the development of PE.
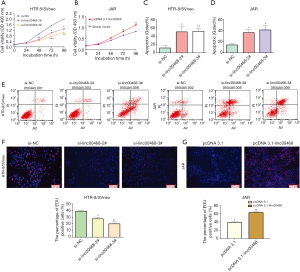
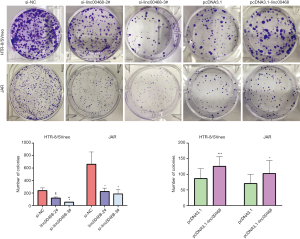
The effect of Linc00468 on cell metastasis and invasion in vitro
Two cell lines were transfected by si-linc00468 and plasmid separately to eliminate the potential role of linc00468 in trophoblast cells. Compared to the si-normal control (siNC) treatment, down-regulation of linc00468 with siRNA #2, and #3 markedly elevated the migration and invasion of the cells (Figure 3A,B). By contrast, linc00468-overexpression impeded the number of trophoblast cells (Figure 3C,D). Flow cytometry analysis showed no significant difference at the G1-G0 phase, either in the inhibition or the overexpression group (Figure 3E).
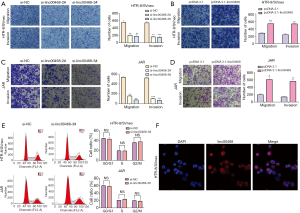
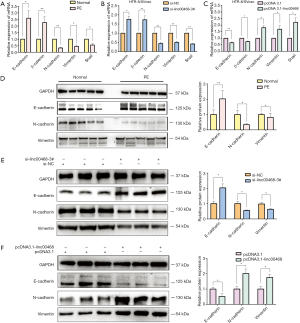
Linc00468 promotes PE progression via regulating EMT
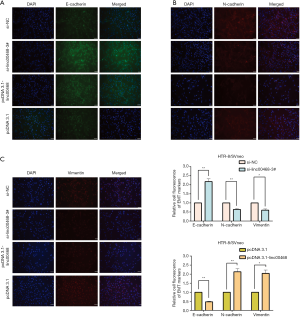
Our previous studies showed that linc00468 contributed to the invasive and migrant ability of HTR-8/Svneo and JAR cell lines. In recent years, the underlying mechanism of EMT has been explored through testing the initial epithelial state or the terminal mesenchymal state in PE, which is affiliated with impaired remodeling. To investigate whether linc00468 influenced the expression of EMT markers in PE, we first used FISH analysis to detect the location of linc00468 in HTR-8/SVneo cells. The results showed that linc00468 was located in both the nucleus and the cytoplasm (Figure 3F). Furthermore, using qRT-PCR, we demonstrated that the PE group induced the expression of E-cadherin and β-catenin and reduced the expression of N-cadherin, Vimentin, and Snail, compared to the healthy group (Figure 4A). Simultaneously, the expression of these markers using qRT-PCR in HTR-8/SVneo cells with upregulated and downregulated linc00468 showed the same results (Figure 4B,C). These data demonstrated that linc00468 effected EMT-related proteins in PE tissues (Figure 4D). Additionally, Western blot analysis showed that the inhibition of linc00468 elevated protein expression of the epithelial marker E-cadherin and suppressed protein expression of the mesenchymal markers N-cadherin and Vimentin, while high levels of linc00468 had the opposite effect (Figure 4E,F). Moreover, we examined the same EMT-associated indicators by immunofluorescence to further confirm whether linc00468 was involved in PE progression by inducing the EMT in the HTR-8/Svneo cell line, which demonstrated the same phenomenon (Figure 5A,B,C,D).
Discussion
Although the molecules and signaling pathways by which PE develops are yet to be elucidated, great strides have been made in PE research. Over recent decades, a myriad of lncRNA functional assays in PE have gained attention and illustrated that lncRNAs play a critical regulatory part in the progression of PE (20). It is thought that pre-eclampsia is a pregnancy-related disorder of angiodysplasia. Some studies support the theory of “superficial trophoblast implantation” (21), because inadequacy or the failure of trophoblast cell invasion, migration, and abnormal extravillous cytotrophoblast (EVT) formation in the uterine wall causes impaired placentation in the development of the placental vascular bed (22,23). In addition, Ferretti et al. provided evidence to corroborate the similar phenotypic properties between cancer and trophoblastic cells in molecular circuits by the disorder of the process of extensive proliferation, migration and invasion (24).
In this study, we showed for the first time that linc00468, a 903-bp lncRNA located on chromosome 6q26 (19), serves a vital role in trophoblast invasion and migration. It was ubiquitously downregulated in the trophoblastic cells and placentas of PE patients, which further explains that the inhibition of linc00468 might result in the development of PE. EDU, CCK8 and clone assays also demonstrated that linc00468 impair the viability of EVTs. Transwell assays revealed that linc00468 has been shown to promote invasion and migration by inducing EMT in vitro. Flow cytometry analysis demonstrated that the suppression of linc00468 promoted apoptosis in cell lines, whereas it did not change the cell cycle. We also showed that overexpressed linc00468 produced the opposite results. Taken together, all the biological results illustrated that lin00468 serves a key role in trophoblast cells invasion and migration.
A follow-up study using FISH analysis showed that linc00468 was localized in both the nucleus and the cytoplasm, indicating its possible involvement in transcriptional and post-transcriptional regulation. To detect the role of linc00468 in regulating the EMT, EMT-relating transcription regulators were detected by qRT-PCR and western blot. Intriguingly, results showed that silencing linc00468 led to the up-regulation of E-cadherin and gradual down-regulation of N-cadherin, Vimentin and Snail; the overexpression of linc00468 led to the up-regulation of N-cadherin, Vimentin, and Snail and the gradual down-regulation of E-cadherin. We also found similar results using an immunofluorescence assay. Through activation of EMT transcription factors (EMT-TFs), we can conclude from the above results that the loss of expression of linc00468 may be directly or indirectly associated with a reduction in the invasive capacity of PE placentas and human cell lines resulting in PE pathogenesis. On the basis of the above description, altered lin00468 is a promising marker of the invasion of EVTs inducing by EMT in PE.
The mesenchymal-like EVT undergo the process of EMT which is the first step from an anchorage-dependent epithelial phenotype (5). Then, EVTs can invade the maternal decidua, remodeling the spiral arteries. As a result, this caused the failure of placentation. Several groups have reported that the EMT is characterized by a pathological dynamic process, where mesenchymal cells replace the organized epithelial phenotype, forming a fibroblast cell phenotype with migratory and invasive properties and lose the intercellular contacts and cellular polarity (25). A previous study confirmed that Vimentin, Snail and N-cadherin exhibited low expression; however, ZO-1 and E-cadherin showed the opposite expression in the villi, decidua and EVT in PE placentas (9). For example, the lncRNA SPRY4-IT1 was recently determined to induce the EMT via the Wnt/β-catenin signaling pathway to cause trophoblast cell dysfunction (26). Additionally, Yu et al. indicated that the silencing of lncMEG3 is associated with the pathogenesis of PE and affects invasion and migration accompanied by increased EMT (27). STAT3, as a vital aspect downstream of RPS4Y1, has been demonstrated to have an important role in regulating the migratory and invasive ability of trophoblasts through the EMT pathway (28). Moreover, other non-coding RNAs, including lnc-ATB (29), microRNA-7 (30), and miR-431 (31) are involved in the EMT in placental trophoblast cells (32).
According to numerous studies, several growth factors and signaling pathways have been implicated in the EMT development, including the TGF-β/SMAD signaling pathway, the WNT/β-catenin signaling pathway, the Notch signaling pathway and others (33), as well as transcription factors, like SNAIL, ZEB and Twist (34).
Future studies are required to investigate the mechanisms underlying inc00468-induced trophoblast invasion in PE. In the future, we plan to study the effects of linc00468 further via the use of primary cells and animal models.
In summary, linc00468 depletion suppressed trophoblast cell migration and invasion and affected the EMT. Our findings offer novel avenues to advance a more detailed knowledge of PE, and suggest the potential of linc00468 as a therapeutic target for lncRNA-based PE therapy.
Acknowledgments
Funding: The study was supported by grants from the National Natural Science Foundation of China (No. 81771603 and No.81801472), the Natural Science Foundation of Jiangsu Province (project numbers: BK20171502, BK20161061, and BK20181080), Jiangsu Province Women and Children Health Scientific Research Project (FRC201755) and the National Key Research Plan of China (grant 2018YFC1002205).
Footnote
Conflicts of Interest: DW serves as an unpaid section editor of Annals of Translational Medicine from Oct 2019 to Sep 2020. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was approved by the Ethics Committee of the First Affiliated Hospital of Nanjing Medical University (2018-SR-252).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Genest DS, Falcao S, Gutkowska J, et al. Impact of exercise training on preeclampsia: potential preventive mechanisms. Hypertension 2012;60:1104-9. [Crossref] [PubMed]
- Noris M, Perico N, Remuzzi G. Mechanisms of disease: Pre-eclampsia. Nat Clin Pract Nephrol 2005;1:98-114. [Crossref] [PubMed]
- Cnossen JS, Vollebregt KC, de Vrieze N, et al. Accuracy of mean arterial pressure and blood pressure measurements in predicting pre-eclampsia: systematic review and meta-analysis. BMJ 2008;336:1117-20. [Crossref] [PubMed]
- Bharadwaj S, Bhat VB, Vickneswaran V, et al. Oxidative stress in preeclamptic mother - newborn dyads and its correlation with early neonatal outcome - a case control study. J Matern Fetal Neonatal Med 2018;31:1548-53. [Crossref] [PubMed]
- DaSilva-Arnold SC, Zamudio S, Al-Khan A, et al. Human trophoblast epithelial-mesenchymal transition in abnormally invasive placenta. Biol Reprod 2018;99:409-21. [Crossref] [PubMed]
- Gluba-Brzozka A, Franczyk B, Bartnicki P, et al. Lipoprotein Subfractions, Uric Acid and Cardiovascular Risk in End-Stage Renal Disease (ESRD) Patients. Curr Vasc Pharmacol 2017;15:123-34. [Crossref] [PubMed]
- de Groot CJ, O'Brien TJ, Taylor RN. Biochemical evidence of impaired trophoblastic invasion of decidual stroma in women destined to have preeclampsia. Am J Obstet Gynecol 1996;175:24-9. [Crossref] [PubMed]
- Brosens I, Renaer M. On the pathogenesis of placental infarcts in pre-eclampsia. J Obstet Gynaecol Br Commonw 1972;79:794-9. [Crossref] [PubMed]
- E Davies J, Pollheimer J, Yong HE, et al. Epithelial-mesenchymal transition during extravillous trophoblast differentiation. Cell Adh Migr 2016;10:310-21. [Crossref] [PubMed]
- Perl AK, Wilgenbus P, Dahl U, et al. A causal role for E-cadherin in the transition from adenoma to carcinoma. Nature 1998;392:190-3. [Crossref] [PubMed]
- Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer 2011;10:38. [Crossref] [PubMed]
- Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 2009;23:1494-504. [Crossref] [PubMed]
- He X, He Y, Xi B, et al. LncRNAs expression in preeclampsia placenta reveals the potential role of LncRNAs contributing to preeclampsia pathogenesis. PLoS One 2013;8:e81437. [Crossref] [PubMed]
- Lu L, Hou Z, Li L, et al. Methylation pattern of H19 exon 1 is closely related to preeclampsia and trophoblast abnormalities. Int J Mol Med 2014;34:765-71. [Crossref] [PubMed]
- Xu Y, Lian Y, Zhang Y, et al. The long non-coding RNA PVT1 represses ANGPTL4 transcription through binding with EZH2 in trophoblast cell. J Cell Mol Med 2018;22:1272-82. [PubMed]
- Wu D, Yang N, Xu Y, et al. lncRNA HIF1A Antisense RNA 2 Modulates Trophoblast Cell Invasion and Proliferation through Upregulating PHLDA1 Expression. Mol Ther Nucleic Acids 2019;16:605-15. [Crossref] [PubMed]
- Zou Y, Li S, Wu D, et al. Resveratrol promotes trophoblast invasion in pre-eclampsia by inducing epithelial-mesenchymal transition. J Cell Mol Med 2019;23:2702-10. [Crossref] [PubMed]
- Yang X, Meng T. Long Noncoding RNA in Preeclampsia: Transcriptional Noise or Innovative Indicators? Biomed Res Int 2019;2019:5437621.
- Pedersen SK, Mitchell SM, Graham LD, et al. CAHM, a long non-coding RNA gene hypermethylated in colorectal neoplasia. Epigenetics 2014;9:1071-82. [Crossref] [PubMed]
- Song X, Luo X, Gao Q, et al. Dysregulation of LncRNAs in Placenta and Pathogenesis of Preeclampsia. Curr Drug Targets 2017;18:1165-70. [Crossref] [PubMed]
- Cartwright JE, Kenny LC, Dash PR, et al. Trophoblast invasion of spiral arteries: a novel in vitro model. Placenta 2002;23:232-5. [Crossref] [PubMed]
- Velicky P, Knofler M, Pollheimer J. Function and control of human invasive trophoblast subtypes: Intrinsic vs. maternal control. Cell Adh Migr 2016;10:154-62. [Crossref] [PubMed]
- Meekins JW, Pijnenborg R, Hanssens M, et al. A study of placental bed spiral arteries and trophoblast invasion in normal and severe pre-eclamptic pregnancies. Br J Obstet Gynaecol 1994;101:669-74. [Crossref] [PubMed]
- Ferretti C, Bruni L, Dangles-Marie V, et al. Molecular circuits shared by placental and cancer cells, and their implications in the proliferative, invasive and migratory capacities of trophoblasts. Hum Reprod Update 2007;13:121-41. [Crossref] [PubMed]
- Li C. New Functions of Long Noncoding RNAs during EMT and Tumor Progression. Cancer Res 2019;79:3536-8. [Crossref] [PubMed]
- Zuo Q, Huang S, Zou Y, et al. The Lnc RNA SPRY4-IT1 Modulates Trophoblast Cell Invasion and Migration by Affecting the Epithelial-Mesenchymal Transition. Sci Rep 2016;6:37183. [Crossref] [PubMed]
- Yu L, Kuang LY, He F, et al. The Role and Molecular Mechanism of Long Nocoding RNA-MEG3 in the Pathogenesis of Preeclampsia. Reprod Sci 2018;25:1619-28. [Crossref] [PubMed]
- Chen X, Tong C, Li H, et al. Dysregulated Expression of RPS4Y1 (Ribosomal Protein S4, Y-Linked 1) Impairs STAT3 (Signal Transducer and Activator of Transcription 3) Signaling to Suppress Trophoblast Cell Migration and Invasion in Preeclampsia. Hypertension 2018;71:481-90. [Crossref] [PubMed]
- Liu X, Chen H, Kong W, et al. Down-regulated long non-coding RNA-ATB in preeclampsia and its effect on suppressing migration, proliferation, and tube formation of trophoblast cells. Placenta 2017;49:80-7. [Crossref] [PubMed]
- Shih JC, Lin HH, Hsiao AC, et al. Unveiling the role of microRNA-7 in linking TGF-beta-Smad-mediated epithelial-mesenchymal transition with negative regulation of trophoblast invasion. FASEB J 2019;33:6281-95. [Crossref] [PubMed]
- Yang X, Meng T. MicroRNA-431 affects trophoblast migration and invasion by targeting ZEB1 in preeclampsia. Gene 2019;683:225-32. [Crossref] [PubMed]
- Moradi MT, Rahimi Z, Vaisi-Raygani A. New insight into the role of long non-coding RNAs in the pathogenesis of preeclampsia. Hypertens Pregnancy 2019;38:41-51. [Crossref] [PubMed]
- Dongre A, Weinberg RA. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol 2019;20:69-84. [Crossref] [PubMed]
- Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 2014;15:178-96. [Crossref] [PubMed]


