
Correlation between the expressions of circular RNAs in peripheral venous blood and clinicopathological features in hepatocellular carcinoma
Introduction
Primary liver cancer is the sixth most frequently diagnosed cancer and fourth leading cause of cancer-related death worldwide. Hepatocellular carcinoma (HCC) accounts for most (75–85%) primary liver cancers worldwide (1). At present, the most effective therapies for HCC are surgical resection and liver transplantation (2). However, HCC patients show a high rate of postoperative tumor recurrence, resulting in poor overall survival (3,4), and the 10-year survival rate is only 7.2% (5). Patients with early stage HCC can undergo curative treatment with 5-year survival exceeding a half (6).
Microvascular invasion (MVI) is a histopathological feature of microscopic metastasis that indicates invasive behavior of HCC and poor survival outcome (7). MVI can only be determined by histological examination on surgical samples, which tremendously limits the preoperative utility of MVI as a tool for treatment decisions (8). For decades, alpha-fetoprotein (AFP) has been one of the most frequently used biomarkers for HCC surveillance worldwide (9). Prothrombin induced by vitamin K absence or antagonist-II (PIVKA-II) is another serum biomarker used for HCC diagnosis, including HCC at early stages (10). Several studies have suggested that increased PIVKA-II serum levels and its elevated tissue expression are associated with MVI (11-13).
Non-coding RNAs are a heterogeneous group of RNA molecules that do not encode proteins but have key regulatory and structural functions in various processes, including tumorigenesis and metastasis (14-16). Circular RNAs (circRNAs) are single-stranded and covalently closed non-coding RNAs, distinct from common linear RNAs, that lack free caps or poly(A) tails (17). circRNAs are stable and conserved molecules as demonstrated by their long half lives in cells (18). Recent studies have reported that circRNAs interact with microRNAs (miRNAs) by stable complementary binding and serve as miRNA sponges to regulate gene expression (19,20). These properties provide circRNAs with the potential to become ideal diagnostic or prognostic biomarkers in diseases. For example, hsa_circ_0000567, derived from exon 2–6 of SET domain‐containing 3 (SETD3), might be a tumor suppressor and a potential diagnosis biomarker in colorectal cancer (21). FECR1, a circRNA consisting of friend leukemia virus integration 1 (FLI1) exons 4-2-3, acts as an upstream regulator to promote breast cancer cell growth by coordinating the regulation of DNA methylating and demethylating enzymes (22). The circRNA sponge for miRNA-7 (ciRS-7, also named as Cdr1as) was the first circRNA to be studied in HCC (23). Our previous study showed that ciRS-7 was significantly correlated with hepatic MVI and AFP level and suggested ciRS-7 might be an independent factor of MVI (24).
Yao et al. demonstrated that circRNA ZKSCAN1 (circZKSCAN1), a zinc finger family gene, was closely with tumor growth, migration, and invasion of HCC (25). Han et al. reported that circRNA circMTO1 (mitochondrial translation optimization 1 homologue) may act as the sponge for oncogenic miR-9 to suppress HCC development and its expression is closely associated with the prognosis of HCC patients, indicating that circMTO1 is a potential prognosis predictor for patient survival (26). Yu et al. identified the circRNA cSMARCA5, which is derived from exons 15 and 16 of the SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 5 (SMARCA5) gene, as the sponge of miR-17-3p and miR-181b-5p to upregulate tissue inhibitor of metalloproteinase-3 (TIMP3). Furthermore, cSMARCA5 was significantly related to tumor differentiation, tumor stage, tumor size and MVI and suppressed the proliferation and migration of HCC cells in vitro and in vivo, making it a potential therapeutic target (27). Zhang et al. reported that plasma cSMARCA5 could be a good diagnostic marker for HCC patients and plasma cSMARCA5 showed improved accuracy when combined with serum AFP levels, with sensitivity, specificity and the area under the receiver operating characteristic (ROC) curve (AUC) of 87.5%, 94.2% and 0.970, respectively (9). Cullin2 (Cul2) circRNA (circ-10720) sponges miRNAs targeting vimentin to promote epithelial-mesenchymal transition in HCC, thereby playing a vital role in tumor metastasis (28). Chen et al. showed that circRNA homeodomain interacting protein kinase 3 (circHIPK3) promotes cell proliferation and migration through aquaporin 3 (AQP3) by sponging miR-124 (29). We recently found that circSETD3 is a novel tumor suppressor of HCC through the circSETD3/miR-421/MAPK14 pathway in vitro and vivo (30).
Blood-based assays are a noninvasive, easily performed and early detection screening tool that can detect circulating tumor cells that originate from the primary tumor site, tumor-associated inflammatory cells and tumor-associated cytokines (31). Identification of noninvasive screening methods for early diagnosis, prediction of MVI, and reduction of disease incidence and mortality for cancer patients is critical. Its accessibility is also because of higher compliance by the HCC patients, high feasibility, reduction of cost, etc. Therefore, a useful preoperative blood-based method to determine depth of tumor infiltration and predict MVI would greatly guide treatment strategy, reduce the early recurrence rate and improve cancer patient prognosis.
In this study, we searched the literature and screened seven circRNAs that have been reported to have biological functions in HCC. The preoperative expression of the seven circRNAs in peripheral venous blood of 88 HCC patients was examined, and correlations between expressions of the circRNAs and clinicopathological characteristics were analyzed. We also examined the correlations of circRNA expression between peripheral venous blood and paired HCC tissues in 48 randomly selected patients. Furthermore, we compared circRNA levels before surgery and three days after surgery. Finally, we analyzed the venous blood level of circSETD3 in HCC patients to determine its predictive value for MVI and performed in vitro assays to evaluate the role of circSETD3 in HCC metastasis.
Methods
Patients and healthy controls
A total of 88 consecutive HCC patients who were confirmed by pathology based on a surgical specimen of the tumor were recruited from West China Hospital from November 2017 to September 2018. All patients were newly diagnosed with HCC and did not receive preoperative therapies, including chemotherapy, radiotherapy or radiofrequency ablation. Detailed clinicopathological data of all patients were extracted from the hospital digital health care system of West China Hospital and are displayed in Table S1. We also recruited 23 healthy controls from the Health Management Center of West China Hospital. All study protocols were approved by the Biomedical Ethics Committee of West China Hospital. The study conformed with the code of ethics of the World Medical Association (Declaration of Helsinki). All participants provided signed informed consent.
Inclusion and exclusion criteria
Patients with HCC who were included in this study met the following criteria: (I) preoperative diagnosis was HCC with no extrahepatic metastasis and no other tumors; (II) postoperative pathology confirmed primary HCC; (III) radical surgical resection was performed for the first time, and no chemotherapy, radiotherapy or other treatments were given; (IV) the patient was over 18 years old, with no limitation on gender, has no obvious mental disorder, and has the right of independent informed consent; (V) the liver function was of Child-Pugh A-B, with no obvious abnormalities of heart, lung and kidney functions, no history of drug abuse or familial genetic diseases, and no metabolic, endocrine, blood system diseases or other serious physical and mental diseases; and (VI) complete preoperative and postoperative clinical data were available. If any of the above criteria was not satisfied, the patient was excluded.
Screening of circRNAs
A systematic search for reports published up to March 1st, 2018 was performed using PubMed, Web of Science and Google. Searches included the keywords “circular RNA”, “hepatocellular carcinoma” and “liver cancer”. To identify all circRNAs, the reference lists of review articles were hand searched to identify further reports. CircSETD3 was verified as a tumor suppressor of HCC in our previous study (30) and was thus included in this study.
Collection of peripheral blood samples and HCC tissues
Peripheral venous blood samples (at least 2 mL) were harvested from the median cubital vein with an ethylenediamine tetraacetic acid anti-coagulated vacutainer and stored at −80 °C until use. All tumor samples used in our study were provided by the Clinical Sample Bank of West China Hospital. The tissues were collected within 15 min after hepatectomy and stored at −80 °C until use.
Total RNA extraction
Total RNA was extracted from tumor samples using TRIzol reagent (Invitrogen, Life Technologies Inc., Darmstadt, Germany) and from 400 µL blood using TRIzol LS reagent (Invitrogen) according to the manufacturer’s instruction. The concentration and quality of the RNA were assessed by measuring absorbance ratios of A260/A280 and A260/A280 using a ScanDrop Nuclear Acid Analyzer (Analytik Jena, Germany). RNA samples with concentrations over 100 ng/µL were considered suitable for experiments.
Reverse transcription (RT) and quantitative polymerase chain reaction (qRT-PCR)
Complementary DNAs (cDNAs) for circRNA and miRNA measurements were synthesized using random and stem-loop primers, respectively. RT reaction was performed using the RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific, USA), and qRT-PCR was performed on each sample in triplicate using Maxima SYBR Green qPCR Master Mix (Thermo Scientific, USA) on the CFX connect real-time system (Bio-Rad, USA). The glyceraldehyde 3-phosphate dehydrogenase (GAPDH) gene was used as an internal control for circRNAs and U6 was used as a control for miRNA-421 detection. Relative expression levels of each gene were calculated using the 2−ΔΔCt method. The ΔCt value was determined by subtracting the Ct value of GAPDH mRNA or U6 from the Ct value of the circRNAs or miR-421. The primers used in qRT-PCR are shown in Table S1.
Cell culture
The Huh7 cell line was purchased from the Cell Bank of Type Culture Collection (Chinese Academy of Sciences, Shanghai, China). Cells were cultured in DMEM/high glucose medium (Hyclone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS) (PAN-Biotek, Aidenbach, Bavaria) and 1% penicillin-streptomycin (Hyclone) in a humidified atmosphere at 37 °C containing 5% CO2. The authenticity of the cell line was verified by DNA fingerprinting before use.
Small interfering RNAs (siRNAs)
SiRNA targeting the junction region of the circSETD3 sequence was purchased from RiboBio (Guangzhou, China). The sequence of circSETD3 siRNA was 5'-CAUCCAGUCAGAAAAAUGGdTdT-3'. Cells were transfected with siRNA oligonucleotides using GenMute Reagent (SignaGen Laboratories, USA) according to the manufacturer’s instructions. Experiments were performed 48 h after transfection.
Transwell assays
Cells were collected, suspended in 500 µL serum-free DMEM medium, and transferred into the upper chamber of 8 µm pore size Transwell 24-well chambers (Merck Millipore, Germany) coated with Matrigel (BD Biosciences, USA). In the lower chamber, 10% FBS-containing medium served as the chemoattractant. After 48 h of incubation, the cells that did not invade through Transwell chamber were removed. The inserts were stained with 0.2% crystal violet and imaged, and cells were counted under an inverted microscope.
Wound healing assay
Cells plated in 6-well plates were transfected as indicated and cultured for 48 h, until they achieved 90% confluence. The cell monolayer was scratched using 10 µL sterile tip and floating cells were removed. The wound was photographed under a microscope after 0, 24 and 48 h.
Statistical analysis
Statistical analyses were carried out with GraphPad Prism version 5.0 (GraphPad Software, San Diego, CA, USA) and SPSS version 22.0 (SPSS Inc., Chicago, IL, USA). Normally distributed data are presented as mean ± standard deviation (SD). P<0.05 was considered statistically significant if the 95% confidence interval (CI) did not include the value 1. Student’s t-test was used where the normality test passed. Risk factors associated with MVI were identified by univariate and multivariate binary logistic regression analyses. The ROC curve was constructed to assess the predictive value of circSETD3 level in peripheral blood samples for MVI. The combined predictive value of circSETD3 expression and PIVKA-II level was analyzed using binary logistic regression. The relationships between the expressions of the seven circRNAs in blood and matched HCC tissues were calculated with Pearson’s correlation test.
Results
Characteristics of patients with HCC and healthy controls
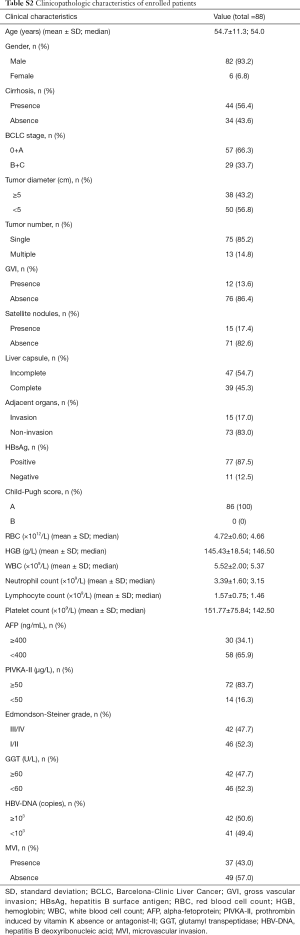
The clinical demographics and tumor characteristics of the 88 HCC patients enrolled in our study are displayed in Table S2. The mean ± SD age of the patients was 54.7±11.3 years (range, 17–81 years), and 93.2% were male. Among the HCC patients, 87.5% were hepatitis B surface antigen (HBsAg)-positive and 56.4% had liver cirrhosis. Patients with AFP over 400 ng/mL and PIVKA-II over 50 µg/L accounted for 34.1% and 83.7%, respectively, of the total patient group. All patients had normal hepatic function (Child-Pugh score A). The Barcelona Clinic Liver Cancer staging system was used to determine the tumor stage. The proportion of patients with stage 0+A tumors was 66.3%. Patients and healthy controls were all of Han nationality and matched in gender, age and ethnicity.
Relationship between the expressions of seven circRNAs and clinical characteristics of HCC
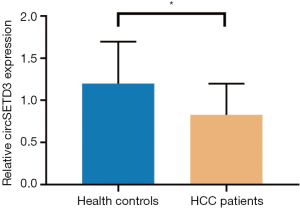
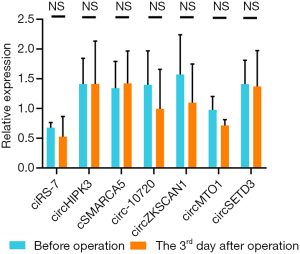
The expressions of all seven circRNAs were measured in peripheral venous blood samples. The frequency distribution and normal P-P plot showed that the expressions of the seven circRNAs were normally distributed (Figure 1). Venous circSETD3 level was significantly reduced in HCC patients compared with levels in healthy controls (P<0.05) (Figure 2), while the other six circRNAs showed no significant differences between groups. We also evaluated levels of the seven circRNAs on the third day after surgery and found that none of circRNAs showed significant changes in expression compared with levels before surgery (Figure 3).
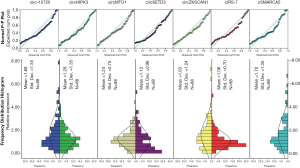
We next analyzed the relationship between the expression of seven circRNAs and HCC patient characteristics (Table 1). We found that all seven circRNAs were not associated with red blood cell count (RBC), white blood cell count (WBC), platelet to lymphocyte ratio (PLR) or neutrophil to lymphocyte ratio (NLR). In addition, the expressions of circHIPK3 and circMTO1 were significantly low in male patients (P=0.042, P=0.031, respectively). Downregulated circSETD3 level was significantly associated with the presence of MVI (P=0.023), the presence of gross vascular invasion (GVI) (P=0.002), PIVKA-II level of 50 µg/L or greater (P=0.006) and incomplete liver capsule (P=0.029). Low expression of circ-10720 was significantly associated with the presence of GVI (P=0.001), and reduced expression of circMTO1 was significantly related to the presence of GVI (P=0.003) and AFP level of 400 ng/mL or greater (P=0.001). Decreased ciRS-7 level was significantly associated with the presence of satellite nodules (P=0.005), and decreased cSMARCA5 level was significantly associated with AFP level of 400 ng/mL or greater (P=0.034). There were no significant correlations between circZKSCAN1 level and clinicopathological parameters.
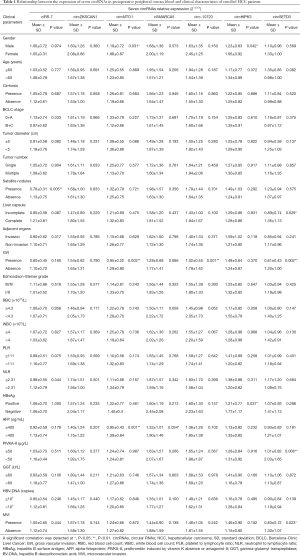
Full table
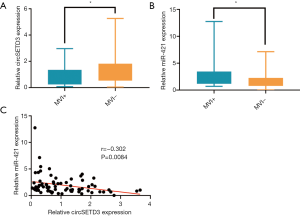
Notably, only down-regulated circSETD3 level in peripheral venous blood samples showed a significant association with the presence of MVI (Figure 4A). Our previous study found that circSETD3 acts as a sponge for miRNA-421 to inhibit HCC cell growth (30). Therefore, we also examined the expression level of miR-421 in venous samples. We found that up-regulated miR-421 level was significantly correlated with the presence of MVI (Figure 4B). Furthermore, venous miR-421 levels were inversely correlated with venous circSETD3 levels (Figure 4C).
Correlations of the levels of the seven circRNAs in peripheral venous blood samples and paired HCC tissues
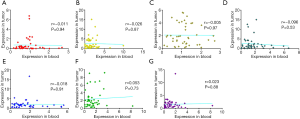
We next analyzed the potential correlation of expression of the seven circRNAs in peripheral venous blood samples and corresponding HCC tissues. However, we found that the expression levels of all seven circRNAs in peripheral venous blood samples were not significantly correlated with levels in paired HCC tissues, as shown in Figure S1.
Venous circSETD3 level as an independent risk factor of MVI
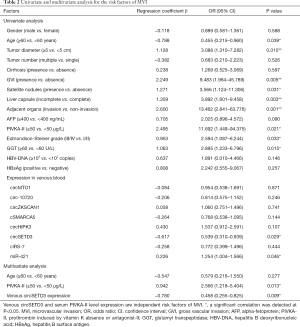
Previous studies reported that the presence of MVI indicates invasive behavior of HCC and is associated with poor survival outcome. We then explored potential risk factors for MVI. As shown in Table 2, univariate analysis revealed that age <60 years, tumor size ≥5 cm, presence of GVI, presence of satellite nodules, incomplete liver capsule, invasion of adjacent organs, PIVKA-II ≥50 µg/L, Edmondson-Steiner grade III/IV, gamma-glutamyl transpeptidase (GGT) ≥60 U/L, upregulated miR-421 and downregulated circSETD3 level were risk factors of MVI. Moreover, a multivariate logistic model showed that low venous circSETD3 expression and serum PIVKA-II level were independent risk factors for MVI.
Full table
Potential predictive ability of venous circSETD3 and serum PIVKA-II levels for MVI
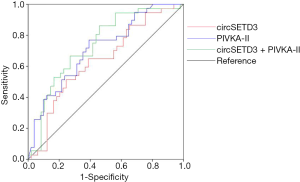
We established ROC curves to evaluate the potential predictive value of the independent risk factors for MVI (Figure 5). Data on AUC are shown in Table 3. PIVKA-II had higher accuracy for the prediction of MVI (AUC =0.713, 95% CI: 0.607–0.819, P=0.001) than circSETD3 (AUC =0.637, 95% CI: 0.519–0.755, P=0.030). The combined predictive value of these two indicators was analyzed by stepwise logistic regression models: logit(p) = −1.719 − 0.677 × (circSETD3) + 2.377 × (PIVKA-II). Notably, the combination of circSETD3 with PIVKA-II showed the highest accuracy compared with either factor alone (AUC =0.736, 95% CI: 0.628–0.844, P<0.001).

Full table
Knockdown of circSETD3 inhibited HCC cell migration and invasion
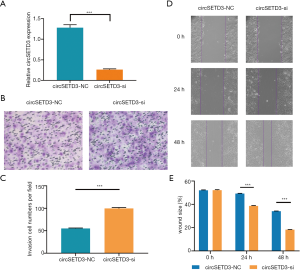
To investigate the effects of circSETD3 knockdown on cell migration and invasion, we next performed wound healing and Transwell assays. Using a backsplice junction-specific siRNA, we successfully knocked down circSETD3 expression in Huh7 cells (Figure 6A). Transwell assays showed that knockdown of circSETD3 strengthened the invasion ability of HCC cells compared with the negative control group (Figure 6B,C). Similarly, downregulation of circSETD3 enhanced migration ability of HCC cells compared with the control group in wound healing assays (Figure 6D,E).
Discussion
In this study, we analyzed the correlations between the preoperative peripheral venous levels of seven circRNAs and clinicopathological characteristics in 88 HCC patients. We also evaluated the ability of venous circRNA levels for preoperative prediction for MVI and compared the accuracy with PIVKA-II, an established biomarker in HCC. Our results suggested that dysregulated venous circRNAs might indicate different tumor characteristics in HCC.
The ability to predict the presence of MVI before surgery could have a major impact on developing therapeutic strategies for patients with HCC. MVI manifests HCC with high ability of invasion and metastasis. The presence of MVI correlates with tumor size, AFP level, histologic grade and high recurrence rate (32,33). Unfortunately, no effective method is available to diagnose MVI before surgery, and MVI is currently only identified by histologic examination of resected specimens obtained from hepatectomy. Therefore, to reduce tumor recurrence rate, it is essential to develop strategies to screen high-risk HCC patients with MVI. Fortunately, because whole blood is composed of a plethora of different cells and a cell-free component, peripheral venous blood circRNAs level is readily available to reflect the overall levels of blood and may be a noninvasive tool to explore tumor biological behavior.
Our results showed that the levels of five circRNAs (circMTO1, circ-10720, cSMARCA5, circHIPK3, and ciRS-7) were significantly associated with one or two clinicopathological characteristics of HCC patients. However, no significant correlation between venous circZKSCAN1 level and clinicopathological parameters was found. Notably, venous circSETD3 level was significantly decreased in the HCC group compared with the healthy control group, suggesting that circSETD3 may function as a cancer suppressor. Importantly, venous circSETD3 level was significantly associated with four clinicopathological characteristics of HCC patients, including serum PIVKA-II level and three highly invasive phenotypes: the presence of MVI, the presence of GVI and incomplete liver capsule. No significant differences were found between preoperative levels of all seven circRNAs and levels on day 3 postoperatively, suggesting high stability and long half-lives of circRNAs in the body.
Consistent with previous findings (24,26,27), lower venous expressions of circMTO1, ciRS-7 and cSMARCA5 indicated HCC with higher malignancy phenotypes, such as presence of satellite nodules and frequent vascular invasion. However, previous studies identified these circRNAs reflecting HCC with higher malignancy in HCC tissues. Although the previous reports suggested a potential strategy for the treatment of HCC patients, the clinical utility of uncovering the expression model in peripheral venous blood of circRNAs must be more valuable. Moreover, many previous studies reported that PLR and NLR may act as prognostic predictors for HCC patients (34-36), as elevated NLR and PLR indicated patients with poorer prognosis. However, we did not find significant correlations between venous circRNA levels and NLR and PLR in this study. We found that decreased venous level of circ-10720 was linked to the presence of GVI. In contrast, Meng et al. found that upregulation of circ-10720 in HCC tissues was associated with aggressive characteristics of HCC (28). The discrepancy between these findings might be due to the different expression pattern of circ-10720 in peripheral venous blood and HCC tissues.
We further explored the relevance of the seven circRNA expressions in peripheral blood by examining their levels in HCC tissue. However, we did not identify a correlation between the expression of the seven circRNAs in venous blood and HCC tissues, suggesting that the venous expression of the seven circRNAs did not directly reflect their expressions in HCC tissue. Notably, however, the abnormal expression of circRNA in venous blood can serve as a risk factor in HCC. In the current study, univariate analysis revealed that age, tumor size, GVI, satellite nodules, liver capsule, invasion of adjacent organs, PIVKA-II, Edmondson-Steiner grade III/IV, GGT, and downregulated circSETD3 level were risk factors for MVI. This result was consistent with previous studies (11,37). Furthermore, multivariable analysis demonstrated that downregulated circSETD3 level and serum PIVKA-II level ≥50 µg/L were independent risk factors of MVI. ROC curve was used to evaluate the predictive ability of venous circSETD3 level for MVI, and the AUC was 0.637 (95% CI: 0.519–0.755, P=0.030) at the cutoff value of 0.545. Notably, combining venous blood levels of circSETD3 and serum levels of PIVKA-II resulted in remarkably improved predictive efficacy for MVI (AUC =0.736, 95% CI: 0.628–0.844, P<0.001), which indicated that peripheral venous circSETD3 may play a vital role in the prediction of hepatic MVI. Taken together, although the expression levels of the seven circRNAs in peripheral venous blood were not significantly correlated with levels in paired HCC tissues, our results showed that preoperative venous levels of circSETD3 could predict MVI. Consistent with these findings, knockdown of circSETD3 promoted HCC cell invasion and migration in vitro. Together these results indicate that circSETD3 may play a major role in the management of patients with HCC.
Although the present study showed that venous circSETD3 level was an independent risk factor of MVI, the mechanism underlying the association between low venous circSETD3 level and MVI has not been clarified. Previous in vitro and in vivo analyses showed that circSETD3 serves as a sponge for miR-421 to inhibit HCC cell proliferation (30). We found the same trend in peripheral blood samples (Figure 3). A single-institution study verified that increased serum PIVKA-II level was a useful and reliable parameter of the presence of MVI in solitary HCC (38). PIVKA-II produced by HCC cells can induce angiogenesis by overexpression of epithelial growth factor receptor (EGFR) and vascular endothelial growth factor (VEGF) and increase the risk of vascular invasion (39).
Our study has several limitations. First, this is a monocentric retrospective study with inherent bias and the number of cases is relatively low. Thus, the current results need to be confirmed in larger external studies. The course of the patients is within one year, and do not reach the follow-up time. Therefore, we cannot analyze the correlation between peripheral venous circRNA expression and prognosis. Second, although the expressions of circRNAs in peripheral whole blood can reflect overall levels, whole blood is composed of a plethora of different cells, including blood cells and the cell-free component. Thus, to reduce confounding bias, a more detailed analysis of circRNA biology in blood components is necessary. In addition, the cell-free component can contain several circRNAs from cells of diverse non-hematopoietic origin. For example, exosome circRNAs secreted from adipocytes can also promote tumor growth (40). Finally, the precise pathways that are regulated by the circRNAs identified in this study have not been confirmed. Therefore, further research is needed to improve our understanding of the role of circRNAs in HCC.
In conclusion, we found that five circRNAs (circMTO1, circ-10720, cSMARCA5, circHIPK3, and ciRS-7) were significantly associated with one or two clinicopathological characteristics (including GVI, but not MVI) of HCC patients, and downregulated venous circSETD3 expression was related with four important clinicopathological characteristics (including MVI) in HCC that are associated with invasion. Importantly, univariate and multivariate analysis identified decreased venous circSETD3 expression as an independent risk factor of MVI in HCC patients. Furthermore, venous circSETD3 expression might play an important role in predicting invasion of HCC especially MVI, and venous circSETD3 level may serve as a potential novel predictive biomarker for HCC invasion.

Full table

Full table
Acknowledgments
We thank Liwen Bianji, Edanz Editing China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript. We gratefully acknowledge JingTao Qiao, Department of Endocrinology and Metabolism, West China Hospital, Sichuan University, Chengdu, Sichuan 610041, China for support. All sources of funding of the study were disclosed.
Funding: This study was supported by the grants of the Key Technology Research and Development Program of the Sichuan Province (No. 2019YFS0208, 2017FZ0082, and 2015SZ0131), National Natural Science Foundation of China (No. 71673193), National Key R&D Program of China (No. 2016YFC1201700), Chinese foundation for hepatitis prevention and control—TianQing Liver Disease Research Fund Subject (No. TQGB20190202), the Natural Science Foundation for Young Scientists and the Science & Technology Planning Project of Gansu Province (No. 18JR3RA058)
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The collection of human specimens was approved by the Biomedical Ethics Committee of West China Hospital (No. 2015202) and the written informed consent were obtained from each patient. All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2008.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424. [Crossref] [PubMed]
- Lee D, Xu IM, Chiu DK, et al. Induction of Oxidative Stress Through Inhibition of Thioredoxin Reductase 1 Is an Effective Therapeutic Approach for Hepatocellular Carcinoma. Hepatology 2019;69:1768-86. [Crossref] [PubMed]
- Bruix J, Gores GJ, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut 2014;63:844-55. [Crossref] [PubMed]
- Hu HT, Wang Z, Huang XW, et al. Ultrasound-based radiomics score: a potential biomarker for the prediction of microvascular invasion in hepatocellular carcinoma. Eur Radiol 2019;29:2890-901. [Crossref] [PubMed]
- Docta RY, Ferronha T, Sanderson JP, et al. Tuning T-Cell Receptor Affinity to Optimize Clinical Risk-Benefit When Targeting Alpha-Fetoprotein-Positive Liver Cancer. Hepatology 2019;69:2061-75. [Crossref] [PubMed]
- Tayob N, Christie I, Richardson P, et al. Validation of the Hepatocellular Carcinoma Early Detection Screening (HES) Algorithm in a Cohort of Veterans With Cirrhosis. Clin Gastroenterol Hepatol 2019;17:1886-1893.e5. [Crossref] [PubMed]
- Li Z, Lei Z, Xia Y, et al. Association of Preoperative Antiviral Treatment With Incidences of Microvascular Invasion and Early Tumor Recurrence in Hepatitis B Virus-Related Hepatocellular Carcinoma. JAMA Surg 2018;153:e182721. [Crossref] [PubMed]
- Lim KC, Chow PK, Allen JC, et al. Microvascular invasion is a better predictor of tumor recurrence and overall survival following surgical resection for hepatocellular carcinoma compared to the Milan criteria. Ann Surg 2011;254:108-13. [Crossref] [PubMed]
- Zhang X, Zhou H, Jing W, et al. The Circular RNA hsa_circ_0001445 Regulates the Proliferation and Migration of Hepatocellular Carcinoma and May Serve as a Diagnostic Biomarker. Dis Markers 2018;2018:3073467.
- Poté N, Cauchy F, Albuquerque M, et al. Performance of PIVKA-II for early hepatocellular carcinoma diagnosis and prediction of microvascular invasion. J Hepatol 2015;62:848-54. [Crossref] [PubMed]
- Kaibori M, Ishizaki M, Matsui K, et al. Predictors of microvascular invasion before hepatectomy for hepatocellular carcinoma. J Surg Oncol 2010;102:462-8. [Crossref] [PubMed]
- Yamashita Y, Tsuijita E, Takeishi K, et al. Predictors for microinvasion of small hepatocellular carcinoma ≤ 2 cm. Ann Surg Oncol 2012;19:2027-34. [Crossref] [PubMed]
- Eguchi S, Takatsuki M, Hidaka M, et al. Predictor for histological microvascular invasion of hepatocellular carcinoma: a lesson from 229 consecutive cases of curative liver resection. World J Surg 2010;34:1034-8. [Crossref] [PubMed]
- Guo H, Ji F, Zhao X, et al. MicroRNA-371a-3p promotes progression of gastric cancer by targeting TOB1. Cancer Lett 2019;443:179-88. [Crossref] [PubMed]
- Wang Y, Liang Y, Yang G, et al. Tetraspanin 1 promotes epithelial-to-mesenchymal transition and metastasis of cholangiocarcinoma via PI3K/AKT signaling. J Exp Clin Cancer Res 2018;37:300. [Crossref] [PubMed]
- Yang Z, Qu CB, Zhang Y, et al. Dysregulation of p53-RBM25-mediated circAMOTL1L biogenesis contributes to prostate cancer progression through the circAMOTL1L-miR-193a-5p-Pcdha pathway. Oncogene 2019;38:2516-32. [Crossref] [PubMed]
- Chen LL. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol 2016;17:205-11. [Crossref] [PubMed]
- Jeck WR, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013;19:141-57. [Crossref] [PubMed]
- Cai H, Zhang P, Xu M, et al. Circular RNA hsa_circ_0000263 participates in cervical cancer development by regulating target gene of miR-150-5p. J Cell Physiol 2019;234:11391-400. [Crossref] [PubMed]
- An Y, Cai H, Zhang Y, et al. circZMYM2 Competed Endogenously with miR-335-5p to Regulate JMJD2C in Pancreatic Cancer. Cell Physiol Biochem 2018;51:2224-36. [Crossref] [PubMed]
- Wang J, Li X, Lu L, et al. Circular RNA hsa_circ_0000567 can be used as a promising diagnostic biomarker for human colorectal cancer. J Clin Lab Anal 2018;32:e22379. [Crossref] [PubMed]
- Chen N, Zhao G, Yan X, et al. A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol 2018;19:218. [Crossref] [PubMed]
- Yu L, Gong X, Sun L, et al. The Circular RNA Cdr1as Act as an Oncogene in Hepatocellular Carcinoma through Targeting miR-7 Expression. PLoS One 2016;11:e0158347. [Crossref] [PubMed]
- Xu L, Zhang M, Zheng X, et al. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol 2017;143:17-27. [Crossref] [PubMed]
- Yao Z, Luo J, Hu K, et al. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol Oncol 2017;11:422-37. [Crossref] [PubMed]
- Han D, Li J, Wang H, et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology 2017;66:1151-64. [Crossref] [PubMed]
- Yu J, Xu QG, Wang ZG, et al. Circular RNA cSMARCA5 inhibits growth and metastasis in hepatocellular carcinoma. J Hepatol 2018;68:1214-27. [Crossref] [PubMed]
- Meng J, Chen S, Han JX, et al. Twist1 Regulates Vimentin through Cul2 Circular RNA to Promote EMT in Hepatocellular Carcinoma. Cancer Res 2018;78:4150-62. [Crossref] [PubMed]
- Chen G, Shi Y, Liu M, et al. circHIPK3 regulates cell proliferation and migration by sponging miR-124 and regulating AQP3 expression in hepatocellular carcinoma. Cell Death Dis 2018;9:175. [Crossref] [PubMed]
- Xu L, Feng X, Hao X, et al. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J Exp Clin Cancer Res 2019;38:98. [Crossref] [PubMed]
- Asgharzadeh S, Salo JA, Ji L, et al. Clinical significance of tumor-associated inflammatory cells in metastatic neuroblastoma. J Clin Oncol 2012;30:3525-32. [Crossref] [PubMed]
- Roayaie S, Blume IN, Thung SN, et al. A system of classifying microvascular invasion to predict outcome after resection in patients with hepatocellular carcinoma. Gastroenterology 2009;137:850-5. [Crossref] [PubMed]
- Shen J, Wen J, Li C, et al. The prognostic value of microvascular invasion in early-intermediate stage hepatocelluar carcinoma: a propensity score matching analysis. BMC Cancer 2018;18:278. [Crossref] [PubMed]
- Zheng J, Cai J, Li H, et al. Neutrophil to Lymphocyte Ratio and Platelet to Lymphocyte Ratio as Prognostic Predictors for Hepatocellular Carcinoma Patients with Various Treatments: a Meta-Analysis and Systematic Review. Cell Physiol Biochem 2017;44:967-81. [Crossref] [PubMed]
- Liao W, Zhang J, Zhu Q, et al. Preoperative Neutrophil-to-Lymphocyte Ratio as a New Prognostic Marker in Hepatocellular Carcinoma after Curative Resection. Transl Oncol 2014;7:248-55. [Crossref] [PubMed]
- Halazun KJ, Hardy MA, Rana AA, et al. Negative impact of neutrophil-lymphocyte ratio on outcome after liver transplantation for hepatocellular carcinoma. Ann Surg 2009;250:141-51. [Crossref] [PubMed]
- Rodriguez-Peralvarez M, Luong TV, Andreana L, et al. A systematic review of microvascular invasion in hepatocellular carcinoma: diagnostic and prognostic variability. Ann Surg Oncol 2013;20:325-39. [Crossref] [PubMed]
- Hirokawa F, Hayashi M, Miyamoto Y, et al. Outcomes and predictors of microvascular invasion of solitary hepatocellular carcinoma. Hepatol Res 2014;44:846-53. [Crossref] [PubMed]
- Fujikawa T, Shiraha H, Ueda N, et al. Des-gamma-carboxyl prothrombin-promoted vascular endothelial cell proliferation and migration. J Biol Chem 2007;282:8741-8. [Crossref] [PubMed]
- Zhang H, Deng T, Ge S, et al. Exosome circRNA secreted from adipocytes promotes the growth of hepatocellular carcinoma by targeting deubiquitination-related USP7. Oncogene 2019;38:2844-59. [Crossref] [PubMed]


