
Performance of modified carbapenem inactivation method and inhibitor-based combined disk test in the detection and distinguishing of carbapenemase producing Enterobacteriaceae
Introduction
Carbapenems, a class of broad spectrum β-lactam antibiotic, have been widely used in clinical practices and play an important role in treatment of multiple drug-resistant bacterial infections, mixed aerobic and anaerobic infections, and infections in immunodeficient patients (1). SENTRY monitoring in United States failed to find Enterobacteriaceae insensitive to carbapenems between 1998 and 2004 (2), carbapenem-resistant Enterobacteriaceae (CRE) has gradually increased around the world, including China, since 2005Jeny (3-6).
The main mechanism underlying the resistance of CRE is the production of carbapenemase (7). Many carbapenems coding genes are located in plasmids, transposon or other removable genetic elements, which promotes the horizontal transfer of resistance genes between gram-negative organisms, and thus it is imperative to develop rapid and accurate methods for the identification of these organisms. The new antimicrobial drug ceftazidime/avibatan can be used to treat carbapenemase-producing Enterobacteriaceae (CPE) which has no metalloenzyme (8), and therefore the detection of genotype is also necessary.
Several approaches have been developed for the detection of carbapenemases in clinical laboratories, such as chromogenic agars, MALDI-TOF MS, Carba NP, CIM, and polymerase chain reaction (PCR) based method (9-11). In these approaches, phenotypic assay is the simplest and easiest to be popularized in primary settings.
The antimicrobial activity may decrease when the carbapenem disk is incubated with CPE, and then the activity of disk is detected to determine whether carbapenemases are produced, which is also known as the modified carbapenem inactivation method (mCIM) (12). The Clinical and Laboratory Standards Institute (CLSI) has recommended combined use of mCIM and EDTA-carbapenem inactivation method (eCIM) to distinguish metallo-carbapenemase from serine-carbapenemase in 20–22 h (13).
Combined disk test (CDT) is another phenotypic assay which was firstly reported for the direct identification of CPE with surveillance rectal swabs in 2013 (14). It is an inhibitor-based test and relies on the fact that metallo-carbapenemase activity can be inhibited by EDTA and serine-carbapenemase be inhibited by phenyl boronic acid (PBA) (15,16). In the present study, the performance of both methods was compared in the detection and distinguishing of carbapenemase production, using PCR as a reference.
Methods
Bacterial isolates
The study was conducted at Renmin Hospital of Wuhan University between January 2017 and December 2018. A total of 101 nonrepetitive carbapenem insensitive Enterobacteriaceae [minimal inhibitory concentration (MIC) ≥2 µg/mL] were collected, including 47 Klebsiella pneumoniae, 26 Escherichia coli, 8 Klebsiella oxytoca, 8 Enterobacter aerogenes,7 Enterobacter cloacae, 2 Citrobacter freundii, 2 Morganella morganii, and 1 Providencia rettgeri. 38 were isolated from urine, 33 from sputum, 12 from blood, 6 from secretions, 4 from bile, 2 from ascites, 2 from pus, 1 from peritoneal dialysate, 1 from cerebrospinal fluid, 1 from feces, and 1 from the tip of a catheter. Isolates were re-examined by MALDI-TOF MS (Bruker Biotyper, Germany). Antimicrobial susceptibility was determined by PhoenixTM 100 (Becton, Dickinson and Company, America). Quality control strains included E. coli ATCC25922, E. coli ATCC35218, K. pneumoniae ATCC BAA-1705, K. pneumoniae ATCC BAA-1706, K. pneumoniae ATCC BAA-2146.
mCIM and eCIM
Detection of CPE was performed with the mCIM and eCIM following the CLSI guideline (13). Two tubes containing 2 mL of Trypticase soy broth (TSB) were used for each strain in parallel: one tube was added with 20 µL of 0.5 M EDTA (Sigma); the other tube had no EDTA. One loop fresh colony of tested strain was taken to each of the tube with 1 µL of inoculating loop. A 10-mg meropenem disk (Oxoid) was incubated in a suspension of tested strain for 4 h at 35 °C. Then, meropenem disks from two tubes were placed on the same Mueller Hinton agar plates, followed by inoculation with the E. coli ATCC 25922 indicator strain. The mCIM showed positive when the inhibition zone diameter was 6–15 mm, or 16–18 mm with small colonies in the inhibitory zone. The eCIM results should only be interpreted if the mCIM result indicated the presence of carbapenemase. A 5-mm increase in the zone diameter for eCIM as compared to that for mCIM suggests the likelihood of produce metallo-carbapenemase.
CDT
As reported by Pournaras et al. (14), 10 µL of 40 mg/mL PBA(Sigma) and 10 µL of 0.5 M EDTA (Sigma) were independently added to 10-mg meropenem disk, which was dried at room temperature. A 10-mg meropenem disk, a meropenem disk plus 10 µL of 40 mg/mL PBA and a meropenem disk plus 0.5 M EDTA were independently pasted on the same Mueller Hinton agar plates which inoculated with 0.5 McFarland of tested isolates, then incubated at 35 °C in ambient air for 16–18 h. If the inhibition zone diameter of meropenem was ≤19 mm and difference in the inhibition zone between meropenem disks with or without inhibitors (PBA, EDTA) was ≥5 mm, producing serine-carbapenemase or metallo-carbapenemase could be defined.
Genotyping of carbapenemase genes
Total DNA was extracted using the boiling method. Carbapenemase genes including blaKPC, blaNDM, blaIMP, blaVIM and blaOXA-48-like genes were amplified using the primers shown in Table 1. Template DNA 1.5 µL, 2× Taq PCR Master Mix (10 µL), and forward and reverse primers (0.5 µL for each) were added to the PCR mixture (final volume: 20 µL), PCR was done as follows: 95 °C for 5 min; 32 cycles of 95 °C for 30 s, 60 °C for 30 s, and 72 °C for 30 s; and 72 °C for 10 min, on the T100™ PCR System (Bio-Rad, America). PCR products were analyzed by agarose gel electrophoresis (SYNGENE, UK).
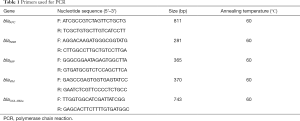
Full table
Statistical analysis
The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), accuracy, and the consistency analysis were determined to assess the performance of mCIM combined eCIM and CDT in the identification and distinguish carbapenemase using PCR results as a standard. Statistical analyses were performed using SPSS 22.0. Cohen’ kappa <0.4, poor consistency; Cohen’ kappa=0.4–0.75, a fair degree of consistency; Cohen’ kappa >0.75, excellent consistency.
Results
Seventy-nine CPEs were detected with PCR in 101 non-repetitive isolates of Enterobacteriaceae, which were insensitive to imipenem or meropenem (MIC ≥2 µg/mL). The CPEs included 38 KPC (38 Klebsiella pneumoniae), 32 NDM (22 Escherichia coli, 3 Enterobacter cloacae, 3 Klebsiella oxytoca, 2 Citrobacter freundii, 1 Klebsiella pneumoniae, and 1 Providencia rettgeri), 5 IMP (3 Klebsiella oxytoca, 2 Klebsiella pneumoniae), 1 VIM (one Enterobacter cloacae), 2 KPC + NDM (2 Klebsiella oxytoca) and 1 NDM + IMP (1 Klebsiella pneumoniae).
In the present study, the coincidence rate of mCIM combined eCIM and CDT was 97.5% (77/79) and 96.2% (76/79) respectively, compared with gene detection. Most of carbapenemase producers identified by two methods were KPC- and NDM-producers, mainly in Klebsiella pneumoniae and Escherichia coli. However, it was worth noting that, one blaIMP-producing isolate of Klebsiella oxytoca had misclassified as serine-carbapenemase using CDT. And two blaKPC+NDM-producing isolates of Klebsiella oxytoca had also misclassified, mCIM combined eCIM showed serine-carbapenemase, while CDT showed metallo-carbapenemase. Of 101 isolates, none of blaOXA-48like was detected, and thus it was unable to show the capacity of both methods in the detection and distinguishing of organisms carrying this gene. The results of mCIM, eCIM and CDT in different isolates are shown in Table 2. Results of detection of control isolates with or without carbapenemase genes were consistent with the expectations for phenotypic tests.
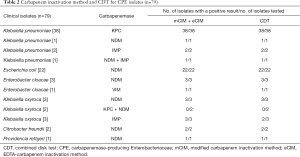
Full table
There was a high agreement between mCIM and PCR assays in the detection of carbapenemase enzymes, except one false-positive on mCIM, which did not identify the five main carbapenemase genes by PCR. The sensitivity was 100%, specificity was 95.5%, к=0.970 (Table 3).
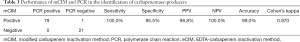
Full table
Two phenotypic methods had similar performance in distinguishing serine-carbapenemase and metallo-carbapenemase. The sensitivity, specificity, PPV, NPV, accuracy, and Cohen’ kappa value are shown in Table 4 respectively.
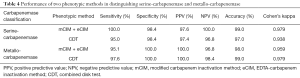
Full table
Discussion
The emergence of CPE has been a global challenge and a major threat to public health worldwide (17). The prevalence of CPE varies among different countries. A recent prospective multinational study investigated the epidemiology of carbapenem-producing Klebsiella pneumoniae and Escherichia coli in Europe (18), and results showed the distribution of four main types of carbapenems (blaKPC, blaNDM, blaIMP and blaOXA-48-like genes) in Klebsiella pneumoniae and Escherichia coli was significantly different among countries: blaOXA-48-like gene was common, and blaVIM was the most uncommon gene. A report from China CRE Network shows that 89% of clinical isolates (n=155) produces carbapenemases, with a majority of isolates producing KPC (50%) or NDM (33.5%)-type gene among K. pneumoniae and E. coli (19). In our study, five major gene types were detected and results showed blaKPC and blaNDM were most common, followed by blaIMP, blaKPC + NDM, blaVIM and blaNDM +IMP. OXA-48-like genes were not identified in the present study.
Our study showed, after overnight incubation, mCIM had favorable performance in the phenotypic detection of carbapenemase producers except one Enterobacter aerogenes showed “false-positive”. It maybe not a false-positive, because of other serine-carbapenemase had not been detected in our study, such as IMI, SME, NMC, GES or other unknown carbapenemase. It just the advantage of phenotypic methods, which was much easier to find unknown resistance genes than target PCR.
The performance of mCIM combined eCIM and CDT were compared with PCR in the classification of carbapenems. Results showed the two phenotypic methods had a high agreement with PCR in the detection of a single gene such as 38 Klebsiella pneumoniae carrying blaKPC, 22 Escherichia coli carrying blaNDM, and the detection of the same type gene carried by CPEs, such as one Klebsiella pneumoniae carrying blaNDM + IMP. For different types of carbapenemase, two phenotypic methods were incorrect for two Klebsiella oxytoca which carried blaKPC + NDM. When metallo-carbapenemase was inhibited by EDTA, the co-existence of serine-carbapenemase could also hydrolyze meropenem, make the inhibition zone diameter decreased and showed serine-carbapenemase positive. So CLSI warned that eCIM was not suitable for the isolate carried both serine and MBL carbapenemase, because it can show false negative (13). And in our study, CDT was also can’t distinguish the two types in one isolate, it might be related with the relative amount and effect between the two inhibitors. Nevertheless, it was few that isolate both encoding serine and MBL carbapenemase. Zhou et al. evaluated four phenotypic methods in the detection of Class A and B CPE in Peking Union, Medical College Hospital, only two isolates carried blaKPC + IMP were collected, among 342 CRE in half a year (20).
Both of mCIM combined eCIM and CDT had a high sensitivity and specificity in distinguishing serine-carbapenemase and metallo-carbapenemase (>95%), and had excellent consistency with gene detection results (к>0.9). The results were in accordance with other study evaluating improved methods based on CIM, like CIMplus test (21).
Of note, 21.8% (n=22) of isolates, which were non-carbapenemase-producing carbapenem insensitive Enterobacteriaceae, were negative to the five tested primers. Most of them were Enterobacter cloacae and Enterobacter aerogenes. Maybe it contained other carbapenemases gene, such as blaGES, blaSME, blaOXA-181, blaOXA-244, or ESBL and AmpC gene over-expression, or loss/alteration of OmpF and OmpC genes (22).
In this study, other carbapenem genes such as blaOXA were not detected simultaneously. Since OXA-48 and related variants had no specific inhibitors available, the CDT was not suitable for detecting OXA-like gene positive isolate (23). At the same time, due to the weak hydrolytic ability to carbapenems, the CIM sometimes fails to detect OXA-48 in a limited time, while the sensitivity of mCIM was improved without compromising from the specificity (24).
The tests we evaluated in this study have many advantages, like low cost, and easy to interpretable. However, there are also some limitations. The test genes were not enough and we need great varieties gene types. And there was no statistical difference between the two phenotypic methods, maybe we need a large number of CPE and non-CPE. However, for experimental operations, CDT is simpler and easy to be commercialized, but needs to be further verified.
Acknowledgments
Funding: This work supported by the Special Science and Technology Cooperation Project of Ningxia Hui Autonomous Region Key R&D Program (2018BFG02008) and the National Science and Technology Key Projects on “Major Infectious Diseases such as HIV/AIDS, Viral Hepatitis Prevention and Treatment” (No. 2017ZX10103005).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was approved by the Ethics Committee of Renmin Hospital of Wuhan University (2016K-C025).
References
- Bradley JS, Garau J, Lode H, et al. Carbapenems in clinical practice: a guide to their use in serious infection. Int J Antimicrob Agents 1999;11:93-100. [Crossref] [PubMed]
- Deshpande LM, Jones RN, Fritsche TR, et al. Occurrence and characterization of carbapenemase-producing Enterobacteriaceae: report from the SENTRY Antimicrobial Surveillance Program (2000-2004). Microb Drug Resist 2006;12:223-30. [Crossref] [PubMed]
- Naas T, Nordmann P, Vedel G, et al. Plasmid-mediated carbapenem-hydrolyzing beta-lactamase KPC in a Klebsiella pneumoniae isolate from France. Antimicrob Agents Chemother 2005;49:4423-4. [Crossref] [PubMed]
- Peleg AY, Franklin C, Bell JM, et al. Dissemination of the metallo-beta-lactamase gene blaIMP-4 among gram-negative pathogens in a clinical setting in Australia. Clin Infect Dis 2005;41:1549-56. [Crossref] [PubMed]
- Wang Q, Wang X, Wang J, et al. Phenotypic and Genotypic Characterization of Carbapenem-resistant Enterobacteriaceae: Data From a Longitudinal Large-scale CRE Study in China (2012-2016). Clin Infect Dis 2018;67:S196-205. [Crossref] [PubMed]
- Wei ZQ, Du XX, Yu YS, et al. Plasmid-mediated KPC-2 in a Klebsiella pneumoniae isolate from China. Antimicrob Agents Chemother 2007;51:763-5. [Crossref] [PubMed]
- Patel G, Bonomo RA. Status report on carbapenemases: challenges and prospects. Expert Rev Anti Infect Ther 2011;9:555-70. [Crossref] [PubMed]
- Livermore DM, Meunier D, Hopkins KL, et al. Activity of ceftazidime/avibactam against problem Enterobacteriaceae and Pseudomonas aeruginosa in the UK, 2015-16. J Antimicrob Chemother 2018;73:648-57. [Crossref] [PubMed]
- Hirsch EB, Chang KT, Zucchi PC, et al. An evaluation of multiple phenotypic screening methods for Klebsiella pneumoniae carbapenemase (KPC)-producing Enterobacteriaceae. J Infect Chemother 2014;20:224-7. [Crossref] [PubMed]
- Monteferrante CG, Sultan S, Ten Kate MT, et al. Evaluation of different pretreatment protocols to detect accurately clinical carbapenemase-producing Enterobacteriaceae by MALDI-TOF. J Antimicrob Chemother 2016;71:2856-67. [Crossref] [PubMed]
- Moran Gilad J, Carmeli Y, Schwartz D, et al. Laboratory evaluation of the CHROMagar KPC medium for identification of carbapenem-nonsusceptible Enterobacteriaceae. Diagn Microbiol Infect Dis 2011;70:565-7. [Crossref] [PubMed]
- Pierce VM, Simner PJ, Lonsway DR, et al. Modified Carbapenem Inactivation Method for Phenotypic Detection of Carbapenemase Production among Enterobacteriaceae. J Clin Microbiol 2017;55:2321-33. [Crossref] [PubMed]
- Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing. 28th editions. CLSI supplement M100-S28. Wayne: CLSI, 2018.
- Pournaras S, Zarkotou O, Poulou A, et al. A combined disk test for direct differentiation of carbapenemase-producing enterobacteriaceae in surveillance rectal swabs. J Clin Microbiol 2013;51:2986-90. [Crossref] [PubMed]
- Pasteran F, Mendez T, Guerriero L, et al. Sensitive screening tests for suspected class A carbapenemase production in species of Enterobacteriaceae. J Clin Microbiol 2009;47:1631-9. [Crossref] [PubMed]
- Tsakris A, Kristo I, Poulou A, et al. First occurrence of KPC-2-possessing Klebsiella pneumoniae in a Greek hospital and recommendation for detection with boronic acid disc tests. J Antimicrob Chemother 2008;62:1257-60. [Crossref] [PubMed]
- Tamma PD, Goodman KE, Harris AD, et al. Comparing the Outcomes of Patients With Carbapenemase-Producing and Non-Carbapenemase-Producing Carbapenem-Resistant Enterobacteriaceae Bacteremia. Clin Infect Dis 2017;64:257-64. [Crossref] [PubMed]
- Grundmann H, Glasner C, Albiger B, et al. Occurrence of carbapenemase-producing Klebsiella pneumoniae and Escherichia coli in the European survey of carbapenemase-producing Enterobacteriaceae (EuSCAPE): a prospective, multinational study. Lancet Infect Dis 2017;17:153-63. [Crossref] [PubMed]
- Zhang Y, Wang Q, Yin Y, et al. Epidemiology of Carbapenem-Resistant Enterobacteriaceae Infections: Report from the China CRE Network. Antimicrob Agents Chemother 2018. [Crossref] [PubMed]
- Zhou M, Wang D, Kudinha T, et al. Comparative Evaluation of Four Phenotypic Methods for Detection of Class A and B Carbapenemase-Producing Enterobacteriaceae in China. J Clin Microbiol 2018. [Crossref] [PubMed]
- Caméléna F, Cointe A, Mathy V, et al. Within-a-Day Detection and Rapid Characterization of Carbapenemase by Use of a New Carbapenem Inactivation Method-Based Test, CIMplus. J Clin Microbiol 2018. [Crossref] [PubMed]
- Babouee Flury B, Ellington MJ, Hopkins KL, et al. Association of Novel Nonsynonymous Single Nucleotide Polymorphisms in ampD with Cephalosporin Resistance and Phylogenetic Variations in ampC, ampR, ompF, and ompC in Enterobacter cloacae Isolates That Are Highly Resistant to Carbapenems. Antimicrob Agents Chemother 2016;60:2383-90. [Crossref] [PubMed]
- Aguirre-Quiñonero A, Martínez-Martínez L. Non-molecular detection of carbapenemases in Enterobacteriaceae clinical isolates. J Infect Chemother 2017;23:1-11. [Crossref] [PubMed]
- Tamma PD, Opene BN, Gluck A, et al. Comparison of 11 Phenotypic Assays for Accurate Detection of Carbapenemase-Producing Enterobacteriaceae. J Clin Microbiol 2017;55:1046-55. [Crossref] [PubMed]


