
Identification of combined T-cell and B-cell reactive Echinococcus granulosus 95 antigens for the potential development of a multi-epitope vaccine
Introduction
Cystic echinococcosis (CE), also known as hydatid disease, is a zoonotic parasitic disease caused by larvae of Echinococcus granulosus (EG), which infects both human and animals globally (1,2). EG is a serious threat to the production of livestock and has a severe effect on health of human. At present, there is no effective treatments for echinococcosis. Inducing massive efforts to carry out research in the area of immunology associated with EG infection with the hope of the development of future vaccine in recent years (3,4). Currently, all the proteins that have shown potential with respect to vaccine development, the Eg95 antigen is the most effective (5). Lightowlers first published research pertaining to the use of the Eg95 antigen in a host recombinant antigen vaccine in 1993 (6,7). The potential vaccine, against the larval membrane protein, conferred 95% protection to immunized sheep against the parasite. The reduction rate of Echinococcus was 96%, the cyst mean was 5.8. These results demonstrated that the Eg95 antigen has potential as a vaccine candidate molecule in preventing the infection of Echinococcosis (8).
There are some specialized chemical moieties which have multifarious functions existing at the surface of antigenic molecules. The groups, known as epitopes, can dictate antigen specificity and induce humoral and cellular immune responses (9). Depending on the identification of the antigenic epitope, cells can be divided into T-cell or B-cell reactive epitopes. In order to research the potential of Eg95 as a potential vaccine further, we studied the ability of this epitope both in humoral and cellular immunity by studying the combined T-cell and B-cell reactive epitopes of Eg95. Using bioinformatics technology, not only did we research the potential for improving vaccine candidate efficiency, but also we greatly reduced the consumption of excess resources that required for the elucidation of new potential vaccine targets (10). It is hoped that this study has facilitated elucidation of an optional peptide vaccine that will help to enhance the cross protection of vaccine. One of the key determinants in generating an effective peptide vaccine involves the screening and identification methods. This study developed a phage display system that integrated the DNA sequence of external epitope proteins or peptides to the gene structure of phage coat protein. It meant that the eligible epitope would accompany phage proteins during reassembly, exposing the external proteins or peptides to the phage surface. As one of the most widely used of these systems, the phage display system which this study used is the filamentous phage display system (11-13). When the coding sequences of the antigen epitope are inserted into the gene of phage, the peptides maintain rendered their natural conformation. This means that gene can be expressed, with the corresponding antigen receptor specificity tested for future potential antigen epitope.
Based on the bioinformatics technology analysis, this study obtained Eg95 protein epitope sequences and further analyze T-cell and B-cell epitope for an antigenic element associated with the EG infection process. An advanced filamentous phage display system was used to predict effective T-cell and B-cell reactive epitopes that could be used for identification and screening of potential vaccine peptides. Although further studies are required for further development of a suitable Echinococcosis epitope peptide vaccine, this study aimed to provide some valuable tools and information for the potential development of the echinococcosis vaccine.
Methods
Plasmid, strains, serum and reagents
E. coli DH5α strains were stored in our laboratory; ER2738 E. coli strains, M13KE phage carrier, PCR kit, RT-PCR kit, restriction enzyme KpnI, restriction enzyme EcoR52I bought from Invitrogen companies in the United States; PMD18-T carrier bought from Dalian Takara Company; Plastic recycling kits, plasmid extraction kit bought from Beijing Tiangen Biological Technology Company; the Ex-Taq DNA polymerase, T4 DNA ligase bought from Dalian Baoxin Biological Company, Ampicillin, X-ray Gal, IPTG, PEG8000 bought from Shanghai Sangon Company; the serum samples of patients who had been diagnosed with CE were provided by the First Affiliated Hospital of Xinjiang Medical University; rEg95 serum primary antibody was prepared by the research group in the previous study which obtained from the polyclonal serum by immunizing New Zealand rabbits. The study was approved by the ethics committee of Xinjiang Medical University (ZACUS-201302255011).
The cloning of Eg95 gene and the construction of prokaryotic expression system of Eg95
The template that recombinant prokaryotic expression vector of pET32a/Eg95 was stored in our laboratory, primer was compounded at Invitrogen Company in Shanghai. Sequences showed in the Table 1. Then amplify the genes by PCR, the system was 50 µL: DNA template 2 µL, 10× buffer solution 5 µL, primer F and R 1 µL,10 mM dNTP 5 µL, 25 mM MgCl2 4 µL, Taq polymerase 0.4 µL, add up to 50 µL of ddH2O. The parameters for PCR were set as follows: 94 °C for 5 min; 94 °C for 10 s, 52 °C for 10 s, 72 °C for 10 s, ×30 cycles; 72 °C for 5 min. The product was detected by 2% agarose gel electrophoresis to detect amplification products. Validations were repeated more than three times.

Full table
Using DNA Fragment Purification Kit (BioDev-Tech) purified amplification fragment and inserted the fragment into plasmid vector pMDl8-T by using T-A cloning kit according to the manufacturer’s instructions and delegate the company sequencing and compared with those of sequence Eg95 genes in GenBank. Recombinant plasmid pMDl8-T-Eg95 and expression vector pet-32a (Novagen) double enzyme, agarose gel electrophoresis separation, cut the plastic recycling. The purified gene fragments and linearization pET-32 had a connection and then amplified in E. coli DH5α, after amplification, extracting plasmid sequence again.
The expression, purification of rEg95 and the antiserum preparation
The recombinant plasmid which contained the purpose gene of E. coil DH5α. had been inoculated in LB containing tendency for 0.1 L IPTG, induced protein expression. The purpose protein expressions were detected by 10% SDS-PAGE BioRad gel image analysis system. Use the Ni-NTA affinity chromatography purification to purify the recombinant proteins. The purified rEg95 was used to immunize rabbits with conventional intradermal multipoint for 4 times, and the serum was isolated. The antiserum titer was determined by the double-diffusion immunoassay.
Predicting T-cell and B-cell epitope of Eg95 molecule
Alternative sequence Eg95 genes by comparing BLAST database, further choose the Eg95 gene sequence (GenBank serial number: HM345607) as the study sequence. Log in UCL-CS bioinformatics website (http://bioinf.cs.ucl.ac.uk/psipred/), select PSIPRED V3.3 to predict Eg95 secondary structure, and logon TMHMM Sever 2.0 TMHMM (http://www.cbs.dtu.dk/services/) to analyze the amino acid sequence of Eg95 protein across the membrane. Respectively using BCEpred (http://www.imtech.res.in/raghava/bcepred/) and LEPS (http://leps.cs.ntou.edu.tw/index.php), by selecting the antigenic index, beta corner, flexibility, hydrophilic, polar and surface accessibility parameters prediction project, comprehensive prediction Eg95 antigen B cell epitope. Using SYFPEITHI (http://www.syfpeithi.de/bin/MHCServer.dll/EpitopePrediction.htm), choose the HLA in MHC types-A0201 and HLA-Db, respectively to the person of Eg95 antigen of T cell epitope and mouse T cell epitope prediction. According to 3Dligandsite server site, predicting tertiary structure of Eg95, and then use RasMol Version of Eg95 antigen software tertiary structure model for display and analysis. Synthesizing find contains both Eg95 antigen B cell epitope also contains Eg95 antigen high scores of T cell epitope and amino acid sequence of T cell epitope region in mice, and find the paragraphs T-joint B cell epitope corresponding gene sequences.
The design and synthesis of primers
According to the cDNA sequence of Eg95 (GenBank serial number: HM345607), by the results of epitope prediction. We choose one of three pieces, according to NEB M13KE carrier enzyme site provided by the company, using DNAman software design contains three proteins peptide precursor and flexible peptide coding sequence and enzyme digestion site specific primers (Invitrogen), synthesized by Invitrogen named Eg95-1, Eg95-2 and Eg95-3. Separately adding leader peptide and flexible peptide, KpnI and flexible EcoR52I enzyme loci. Primer sequences are showed in Table 1.
The cloning and identification of T cell and B cell epitope fragment
Each epitope fragments were obtained by PCR method. First Extract the pMDl8-T-Eg95 by a small plasmid DNA extraction kit, then solubled in TE buffer after its concentration were determined by spectrophotometry method. According to characteristics of NEB phage display system provided by the company, designed containing M13 phage PIII protein precursor peptide and flexible peptide coding sequence and enzyme sites primers (Table 1), synthesized by the Shanghai Invitrogen company. PCR volume 50 µL, in addition to the different primers the remaining reagent and gene amplification were consistent. The parameters for PCR were set as follows: 94 °C for 5 min; 52 °C for 30 s, 94 °C for 30 s, and 72 °C for 30 s, ×30 cycles; 72°C for 10 min. Using agarose gel electrophoresis tested the amplification products, amplification products expected size of 110–190 bp. Contains T-A cloning kit (TaKaRa) was used to construct the epitopes of restructuring pMDl8-T, then entrusted company sequencing. All validations were repeated more than three times.
The building and identification of M13KE displaying peptide system
The phage display system kit provided by NEB company. Extracted the right sequence of epitope peptide sequence restructuring pMDl8-T, and the E. coil ER2738 (NEB) amplification of phage M13KE (NEB) DNA by Insert small alkali denaturation method, respectively with EcorI and 52 KpnI double enzyme, using agarose electrophoresis separation enzyme and cut the clip and rubber cutting recovery (13). After cutting enzyme epitope pieces and phage carrier with the appropriate mixed ratio, and T4 ligase (TaKaRa) 16 °C after connection for the night, conversed into the state of bacteria E. coil ER2738. Making E. coil ER2738, IPITG/X-gal cool to about 50 °C, then dumped to LB which contained tetracycline 37 °C cultivate 6 h. After small plaque and develop as a template, using the kit provided by primers (Table 1) after the preliminary appraisal on the PCR sequencing again.
The cultivation and collecting of the recombinant phage
According to the instructions of kit, a small-scale culture system was established using recombinant phage with correct insertion sequence and tetracycline LB liquid. Making 300 mL of the logarithmic growth E. coil ER2738 fresh culture inoculate in LB liquid medium of 30 mL tetracycline, 37 °Cfor 2 h, adding 15 mL recombinant phage cultures, 28 °C for 4 h. Making the recombinant phage cultures centrifuge for 10 min, under 4 °C 10,000 r/min, taking supernatant with centrifugal again, then adding 1/6 volume PEG8000/NaCl to the supernatant setting for the night at 4 °C. The next day making 12,000 r/min, the centrifugal 15 min, then adding 1/30 cultures supernatant of TBS to the precipitation, then 10,000 r/min 15 min to remove the insoluble impurities centrifugal, repeat twice, finally take precipitate to TBS saving at 20 °C. According to the reference of the methods of determination of the purified recombinant phage concentration, the computation formula is as follows: volume of phage (virions)/mL = (A269 A310) ×6×1,016/ phage bases.
Western blot
Western blot analysis methods were used for detections of the recombinant antigen epitope. Use 12%SDS-PAGE separation gel, containing different restructuring PIIIprotein (rPIII) phage on sample quantity are 3×1014, then stained by Coomassie brilliant blue to observe the rPIII expression. After SDS-PAGE making 23V electricity transfer 45 min to PVDF membrane, use TTBS which contained 5% calf serum closed for the night at 4 °C. The purification of Eg95, which were prepared from the recombinant protein immune rabbit polyclonal serum and CE diagnosed serum samples respectively, were diluted with 1:200 as the primary antibodies. The latter dilution with 1:1,000 were used as the second antibodies (the primary antibodies of rabbit polyclonal serum correspondingly add goat anti-rabbit antibody; the primary antibodies of the CE diagnosed patients serum and healthy serum correspondingly add anti-human that secondary antibodies),both incubate 37 °C for 2 h, fully wash then use DAB method to colour. The wild-type phage display system without inserted fragments was set as the control.
ELISA
The polystyrene board with three epitopes were packaged overnight, after been closed and washed, respectively added the rEg95 rabbit antiserum, CE patients’ serum and normal serum. Then the polystyrene board were washed three times and added the fresh diluted enzyme label antibody 0.1 mL each hole, incubate 37 °C for 1 h, added the coloring solution and the terminated liquid in turn. The absorbance values of 450 nm were detected by the micro plate spectrophotometer (blank: the same volume of BSA package; negative control: package without primary antibodies, the diluent instead of the antiserum; positive control: the rEg95 envelope antigen with the same volume).
Results
T-B combined epitope of Eg95 antigen
Comprehensive analysis and contrast of Eg95 antigen protein the secondary structure, the membrane structure and the B cell epitope, ultimately determine three B combined epitopes are 11–29 aa, 51–79 aa and 100–110 aa. Using Eg95 genes as the templates to build T-B combined epitope (Table 2).

Full table
The result of the epitopes peptide segments amplificated by PCR
The pET32a/Eg95 recombinant plasmid was used as the template for PCR amplificated three T-B combined epitopes (Figure 1). The expression of the three recombinant PMD18-T/Eg95-1, PMD18-T/Eg95-2 and PMD18-T/Eg95-3 were identified by the established PCR system. The amplification products were detected by 2% agarose gel electrophoresis, the expected size of each combined epitopes was displayed in Figure 1.
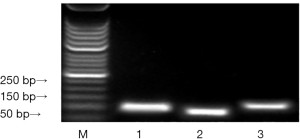

The result of the recombinant phage by PCR
The fuzzy blue plaque from the infected strains were formed under the induction of IPTG. Each clone was Picked to make a small amount of liquid culture, then detected by PCR. The size of wild phage P III proteins gene is 110 bp, amplificational products should be P III proteins gene size add each T-B combined fragment size, Results displayed the same as the expected. The size of Eg95-1, Eg95-2, Eg95-3 were 192, 168 and 222 bp respectively (Figure 2).
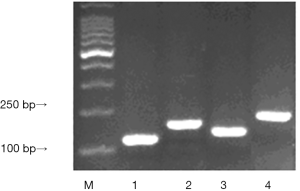
The result of the recombinant phage by SDS-PAGE
M13KE phage PIII actual size is 42.5 kDa, while the band on SDS-PAGE gel position was shown between 60 to 65 kDa. Compared with the wild-type phage, after the insertion of fragments, the molecular sizes of all the three proteins were a little higher, but fewer difference showed between each other, on this PAGE, the result was not obvious (Figure 3).
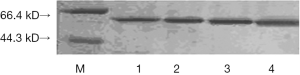

Western blot results
The experiment results (Figure 4) showed that the constructed Eg95 three epitopes can be rEg95 rabbit antiserum, Echinococciasis patients serum recognized and had immune reaction, but according to the result can be found that Eg95-1 had a weaker reaction, Eg95-2 and Eg95-3 had much higher immune reaction than the Eg95-1. It could be inferred that the antigenicity of Eg95-2 and Eg95-3 are stronger than Eg95-1. In our study, the Eg95-2 and Eg95-3 may be antigen epitope.
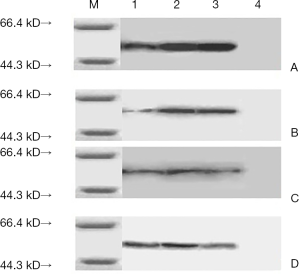
The ELISA results
The test results showed the differences. the antigen-antibody reaction intensity was evaluated by the absorbance value size, the results were showed in Table 3.
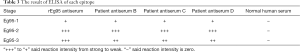

Full table
Discussion
CE is a disease caused by infection with the stage larvae of the parasitic hydatid worm EG. This parasite infects body organs such as the liver and the lungs, causing potentially serious damage to human health (14). In order to ameliorate potential problems caused by the zoonotic parasite, development of an appropriate vaccine to prevent the on set of infection is vital (15-17).
Antigen epitope vaccine shave a vital role in disease prevention. Antigens are immunogenic molecules potentially composed of hundreds of thousands of amino acids. Antigen epitopes, however, contain only a portion of the amino acids present in the antigen itself (18). Generating an effective antigen epitope vaccine requires epitopes that trigger both T-cell and B-cell immune responses (19,20). Through the bioinformatics analyses, this study elucidated three combined T-cell and B-cell reactive epitope nucleotide sequences that were reactive against a larval EG membrane protein. This was facilitated by building a phage display library using the M13KE phage carrier. The application of phage display technology realizes the expression of short peptides on the phage surface, such as the designate epitopes. Potential B-cell and T-cell reactive epitopes could be tested for antigen reactivity with subsequent identification. ELISA and Western blot were used to test the synthesis of the antigen epitopes by the phage display library.
In the early 1990’s, Smith pioneered phage display technology, a successful molecular biotechnology application, allowed the presentation of exogenous peptides on phage surfaces. This technology has been widely applied to scientific research, giving rise to rapid developments in peptide manipulations. The phage display technology realizes rapid genetic manipulations with relatively quick phenotypic responses. Through the phage display system, this study chose the M13KE phage as the carrier for the larval membrane peptide epitopes of the Eg95 antigen. Three separate epitopes were presented by the phage carrier system, which were named as M13KE/Eg95-1, M13KE/Eg95-2 and M13KE/Eg95-3 respectively. After been purified by the PEG/NaCl purification, the expressions of the three T-cell and B-cell reactive epitope peptides were confirmed by SDS-PAGE.
Western blot analysis was subsequently used to assess antisera raised against the three epitope peptides. The three T-cell and B-cell reactive epitopes displayed different degrees of reactivity. The Eg95-1, Eg95-2 and Eg95-3 epitopes were all deemed to have potential suitability as vaccine candidates, the reactivities of which were confirmed by a series of experiments on the rabbit anti rEg95 serum and CE patient serum.
Further analyses were performed to test the potential efficacy of all three short epitopes. The peptide epitopes were tested for resistance against rEg95 rabbit serum. In addition, serum samples which extracted from patients infected with CE were used to test the potential reactivity of all the three epitopes. ELISA test results were performed to directly observe the reactivity of the short peptide sequences. When antiserum was used in resistance tests to assess the efficacy of the epitopes, the three different T-cell and B-cell epitope peptides resulted in different reaction intensities. Absorbance values attained from the analysis demonstrated no cross-reaction between healthy people and CE infected patients, each of the short peptides had adequate specificity. Although the reactivity of the epitope peptide Eg95-1 was weaker in the serum tested, both the Eg95-2 and Eg95-3 exhibited strong reactivity with each of the serum samples analyzed.
The results of the western blot experiments during this study showed that all the three T-cell and B-cell reactive epitope peptides displayed antigenicity. Degree test of immune reactivity associated with each epitope shown that Eg95-1 had a weaker antigenicity. However, both Eg95-2 and Eg95-3 displayed good antigenicity through the test when rabbit anti rEg95 was used and the increased reaction strength were observed. Further could It’s inferred that these two epitope peptides could become to the future vaccine candidates in the prevention of CE.
Acknowledgments
Funding: This work was supported by the National Natural Science Foundation (No. 81760372, 31160194, 81260253).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was approved by the ethics committee of Xinjiang Medical University (ZACUS-201302255011).
References
- Hu D, Song X, Xie Y, et al. Molecular insights into a tetraspanin in the hydatid tapeworm Echinococcus granulosus. Parasit Vectors 2015;8:311. [Crossref] [PubMed]
- Bhutani N, Kajal P. Hepatic echinococcosis: A review. Ann Med Surg (Lond) 2018;36:99-105. [Crossref] [PubMed]
- Zhang F, Li S, Zhu Y, et al. Immunization of mice with egG1Y162-1/2 provides protection against Echinococcus granulosus infection in BALB/c mice. Mol Immunol 2018;94:183-9. [Crossref] [PubMed]
- Tamarozzi F, Akhan O, Cretu CM, et al. Prevalence of abdominal cystic echinococcosis in rural Bulgaria, Romania, and Turkey: a cross-sectional, ultrasound-based, population study from the HERACLES project. Lancet Infect Dis 2018;18:769-78. [Crossref] [PubMed]
- Siles-Lucas M, Casulli A, Conraths FJ, et al. Laboratory Diagnosis of Echinococcus spp. in Human Patients and Infected Animals. Adv Parasitol 2017;96:159-257. [Crossref] [PubMed]
- Liu F, Fan X, Li L, et al. Development of recombinant goatpox virus expressing Echinococcus granulosus EG95 vaccine antigen. J Virol Methods 2018;261:28-33. [Crossref] [PubMed]
- Liu F, Li L, Liu Y, et al. Development of reverse genetics system for small ruminant morbillivirus: Rescuing recombinant virus to express Echinococcus granulosus EG95 antigen. Virus Res 2019;261:50-5. [Crossref] [PubMed]
- Ma X, Zhou X, Zhu Y, et al. The prediction of T- and B-combined epitope and tertiary structure of the Eg95 antigen Echinococcus granulosus. Exp Ther Med 2013;6:657-62. [Crossref] [PubMed]
- Tang ZM, Tang M, Zhao M, et al. A novel linear neutralizing epitope of hepatitis E virus. Vaccine 2015;33:3504-11. [Crossref] [PubMed]
- Larrieu E, Mujica G, Araya D, et al. Pilot field trial of the EG95 vaccine against ovine cystic echinococcosis in Rio Negro, Argentina: 8 years of work. Acta Trop 2019;191:1-7. [Crossref] [PubMed]
- Pourseif MM, Moghaddam G, Saeedi N, et al. Current status and future prospective of vaccine development against Echinococcus granulosus. Biologicals 2018;51:1-11. [Crossref] [PubMed]
- Ebrahimipour M, Afgar A, Barati M, et al. Evaluation of the antigenic epitopes of EgAgB/1 and EgAgB/4 subunit antigens in G1 and G6 genotypes of Echinococcus granulosus using bioinformatics. Gene Reports 2019;15:100361. [Crossref]
- Sotillo J, Toledo R, Mulvenna J, et al. Exploiting Helminth-Host Interactomes through Big Data. Trends Parasitol 2017;33:875-88. [Crossref] [PubMed]
- Conchedda M, Caddori A, Caredda A, et al. Degree of calcification and cyst activity in hepatic cystic echinococcosis in humans. Acta Trop 2018;182:135-43. [Crossref] [PubMed]
- Kinkar L, Laurimäe T, Acosta-Jamett G, et al. Global phylogeography and genetic diversity of the zoonotic tapeworm Echinococcus granulosus sensu stricto genotype G1. Int J Parasitol 2018;48:729-42. [Crossref] [PubMed]
- Huang F, Dang Z, Zhang H, et al. Comparative study on secretome and transmembranome of immature and mature metacestodes of Echinococcus multilocularis. Vet Parasitol 2017;245:153-9. [Crossref] [PubMed]
- Umair S, Pernthaner A, Deng Q, et al. Preliminary evaluation of a thermosensitive chitosan hydrogel for Echinococcus granulosus vaccine delivery. Vet Parasitol 2017;236:117-20. [Crossref] [PubMed]
- Pan W, Chen DS, Lu YJ, et al. Genetic diversity and phylogenetic analysis of EG95 sequences of Echinococcus granulosus: Implications for EG95 vaccine application. Asian Pac J Trop Med 2017;10:524-7. [Crossref] [PubMed]
- Pourseif MM, Moghaddam G, Naghili B, et al. A novel in silico minigene vaccine based on CD4+ T-helper and B-cell epitopes of EG95 isolates for vaccination against cystic echinococcosis. Comput Biol Chem 2018;72:150-63. [Crossref] [PubMed]
- Larrieu E, Poggio TV, Mujica G, et al. Pilot field trial of the EG95 vaccine against ovine cystic echinococcosis in Rio Negro, Argentina: Humoral response to the vaccine. Parasitol Int 2017;66:258-61. [Crossref] [PubMed]


