Evaluation of the association between the −1304T>G polymorphism in the promoter of the MKK4 gene and the risk of colorectal cancer: a PRISMA-compliant meta-analysis
Introduction
The morbidity and mortality of colorectal cancer (CRC) has increased rapidly worldwide (1,2). In China, CRC is the fifth most common cause of tumor death (3). The occurrence of CRC is continuously rising despite the development of comprehensive treatment involving radiotherapy, chemotherapy and surgery (4). Epidemiological studies on CRC revealed multiple influencing factors, such as smoking, alcohol consumption and sex (5). In addition to an unhealthy lifestyle and environmental risk factors, genetic factors also influence the development of CRC. These genetic factors are usually involved in the cellular signal transduction of the mitogen-activated protein kinase (MAPK) pathway, which contributes to apoptosis, inflammation and tumorigenesis (6). Previous studies indicated that genetic risk factors accounted for approximately 35% of the causes of CRC cases (7). Recently, several studies focused on the exploration of the underlying molecular mechanisms regulating CRC carcinogenesis, as well as the crucial roles of genetic factors during the occurrence and development of CRC (8,9). Increasing evidence has indicated that CRC is a disease involving both epidemiological and genetic factors despite the limited understanding of its pathogenesis. Further studies are required to demonstrate the potential relationship between the genetic susceptibility of CRC and key gene mutations.
Single nucleotide polymorphisms (SNPs) are characterized by high density and genetic stability in the genome and are the main source of genetic differences between individuals (10). It is also fundamental for the pathogenesis of many genetic diseases, such as tumors. Moreover, multiple studies confirmed that SNPs as well as age and alcohol consumption were risk factors for the cause of CRC (11-14).
MKK4 is an important component of the MAPK signaling pathway and the central link of the oncogene Ras signaling pathway (15). The polymorphism of the MKK4 gene was reported to affect the efficiency of MKK4 transcription initiation and is associated with the development, progression and prognosis of colon cancer (16-22). Recently, several mutations were detected in exons 4 and 9 of the MKK4 gene in tumor tissues (23,24). However, there were no statistically significant associations between their data and previous results (25). Our present study indicated that mutations in the coding region of MKK4 were not common events in CRC. Based on this result, we hypothesize that CRC with a high incidence in China is associated with the MKK4 −1304T>G polymorphism. We further summarize relevant data from case-control studies. Based on the GenBank dbSNP database, we found 4 common SNPs (rs3809728, rs2190853, rs9892151 and rs3826392), which are located in the promoter region of the MKK4 gene. While other SNPs were in the linkage disequilibrium, rs3826392 was selected as the experimental subject. We conducted a meta-analysis of 4 published case-control studies and quantified the synthetic evidence with strict methods to accurately assess the relationship between the MKK4 −1304T>G polymorphism and the risk of CRC.
Methods
Publication search
We conducted systematic literature searches in the following electronic databases: PubMed, Embase, Cochrane Library, and CNKI, covering all published studies. We searched the databases by the following phrases: “mitogen-activated protein kinase kinase 4”, “MKK4”, “-1304T>G”, “-1304G>T”, “polymorphism”, “colorectal cancer”, “Colorectal Neoplasms”, “Colon cancer” and “rectal cancer”. For example, in the PubMed database, our search strategy was as follows:
#1: (((mitogen-activated protein kinase kinase 4[Title/Abstract]) OR MKK4[Title/Abstract]) OR -1304T>G[Title/Abstract]) OR -1304G>T[Title/Abstract],
#2: ((polymorphism) OR single nucleotide polymorphism) OR SNP[Title/Abstract]),
#3: (((colorectal cancer[Title/Abstract]) OR colorectal neoplasms[Title/Abstract]) OR colon cancer[Title/Abstract]) OR rectal cancer[Title/Abstract],
#4: #1 AND #2 AND #3.
We screened all the research publications and selected all eligible studies. We also examined other related articles in their bibliographies. Only studies published with complete text were included in this study. We only used publications that covered the wider range of information when overlapping articles were found. There were no language restrictions in relevant reports identified.
Inclusion and exclusion criteria
All articles included were required to meet the following criteria: (I) case-control studies of CRC with the MKK4 −1304T>G polymorphism; (II) the study supplied available genotype frequencies in cancer patients and controls; (III) the study provided sufficient published data to estimate an odds ratio (OR) with 95% confidence interval (CI); and (IV) CRC type cancer. The exclusion criteria included the following: (I) the study lacked detailed genotype frequencies; and (II) the article did not conform to the Hardy-Weinberg equilibrium (HWE).
Data extraction
Articles were checked by 2 investigators independently (Rui Bai and Cheng Yuan) to exclude irrelevant and overlapping studies. All authors were involved to discuss the differences. Information was selected in accordance with a protocol: first author’s surname, publication year, ethnicity, source of controls, and the number of patients and controls for each genotype. Factors, such as smoking history, drinking history and sex, were also included. HWE was calculated by the χ2 test based on the distribution of the 2 polymorphic genotypes in the control. P<0.01 was considered a significant imbalance.
Evaluation of study quality
The Newcastle-Ottawa Scale was used to conduct the methodological quality evaluation of the studies for the cohort study. Two reviewers processed the assessments independently, and all authors approved the final decision by consensus.
Statistical analysis
We calculated the OR with 95% CIs to assess the strength of the association between the MKK4 −1304T>G polymorphism and the risk of CRC susceptibility, which were based on the genotype frequencies in cases and controls. The pooled ORs were used for 5 models as follows: allelic model (T vs. G), homozygous model (TT vs. GG), heterozygous model (TG vs. GG), dominant genetic model (TG + TT vs. GG), and recessive model (TT vs. TG + GG). We also used the χ2-based Q statistic test to analyze the heterogeneity between studies. If heterogeneity was considered insignificant (P>0.1), the fixed effects model was used; otherwise, the random effects model based on the Mantel-Haenszel method was applied. All analyses were performed by Review Manager (version 5.3, The Cochrane collaboration).
Results
Characteristics of studies
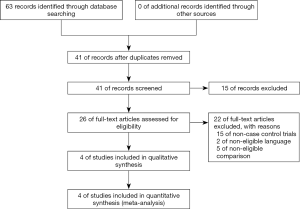
The 1,255 cancer patients and 1,181 control from the included 4 case-control studies met the inclusion criteria (5,26-28). The flow chart shows the paper selection procedure (Figure 1). The main information from the included articles, such as the cases of three genotypes (TT, TG, and GG), ages, sexes, drinking status, smoking status, etc. The baseline characteristics of patients and controls from the included studies are summarized in Table 1.

Full table
Main results
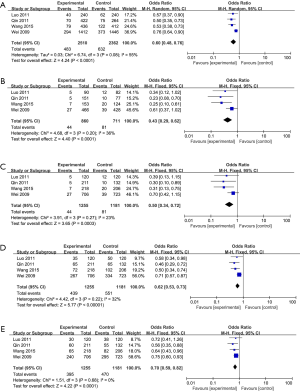
The correlation between the MKK4 −1304T>G polymorphism and the risk of CRC is shown in Figure 2. The results revealed that the MKK4 −1304T>G polymorphism could reduce the risk of CRC (G vs. T: OR, 0.60, 95% CI: 0.48–0.76, P<0.0001, Figure 2A; GG vs. TT: OR, 0.43, 95% CI: 0.29–0.62, P<0.0001, Figure 2B; GG vs. TT + TG: OR, 0.50, 95% CI: 0.34–0.72, P=0.0003, Figure 2C; TG + GG vs. TT: OR, 0.62, 95% CI: 0.53–0.73, P<0.00001, Figure 2D; and TG vs. TT + GG: OR, 0.70, 95% CI: 0.59–0.82, P<0.0001, Figure 2E).
Evaluation of heterogeneity
Except for the allele model (I2=55%, Ph=0.08), there were no significant heterogeneities in any other gene models (GG vs. TT: I2=36%, Ph=0.20, Figure 2B; GG vs. TT + TG: I2=23%, Ph=0.27, Figure 2C; TG + GG vs. TT: I2=32%, Ph=0.22, Figure 2D; and TG vs. TT + GG: I2=0%, Ph=0.68, Figure 2E).
Sensitivity analysis
By using one-way sensitivity analysis, each of the included studies was deleted one-by-one to determine the extent to which individual studies affected the overall OR estimate. The study by Wei et al. (5) was the main source of heterogeneity. When the study was excluded, the heterogeneity significantly decreased (T vs. G: I2=0%, Ph=0.90).
Publication bias
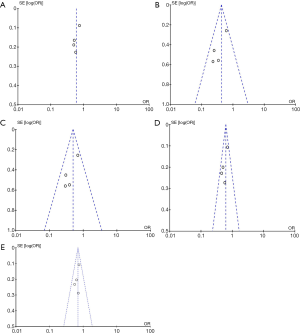
Figure 3 presents the funnel plots for the meta-analysis. Funnel plots did not show any evidence of clear asymmetry.
Discussion
MAPKs are a class of serine/threonine-like protein kinases and play important roles in eukaryotic cells to mediate extracellular signals to intracellular responses (29). All eukaryotic cells express MAPK pathway components, which are MAPK kinase kinase (MKKK), MAPK kinase (MKK) and MAPK (30). These 3 kinases can be activated in sequence and conduct extracellular signals through the tertiary kinase cascade to coregulate cellular physiological and pathological processes, including proliferation, gene expression, differentiation, mitosis, cell survival and apoptosis (31).
MKK4 is a member of the MAPK pathway, whose roles during tumorigenesis are complex (15). In cancer cell lines, MKK4 is a candidate tumor suppressor and is emphasized by its changes (5). Previous studies indicated that JNK and P38 had tumor suppressive effects and that MKK4 was a member of the MAPK family that activated JNK and P38 pathways at the same time (32). MKK4 gene changes, such as homozygous deletions and gene mutations, occur in most tumors. The expression levels of MKK4 were reported to be downregulated in liver cancer, endometrial cancer, lung cancer, and nasopharyngeal carcinoma (33-35). Functional mutations or reduced expression of MKK4 were detected in biliary carcinomas (36). Prostate and gastric cancers have high activation or overexpression of MKK4 (37,38).
Related studies suggested that −1304T>G in the MKK4 promoter region in nasopharyngeal carcinoma was associated with susceptibility to cancer and that the risk of disease decreased as the frequency of the −1304G allele increased (39). Functional −1304G variants in the MKK4 promoter in lung cancer reduced the risk of lung cancer by increasing promoter activity, and G variants were reported as markers of lung cancer susceptibility (34). Heterozygotes and mutants in prostate cancer and breast cancer also showed a low prevalence compared to wild-type (17,38). Our meta-analysis showed that the MKK4 −1304T>G polymorphism was associated with susceptibility to CRC.
It was noteworthy that sensitivity analysis found that research from Wei et al. (5) was the main source of heterogeneity in allele models. However, even so, we should be cautious about this result. First, Wei’s research had a large sample size and was high-quality research. Second, although the I2 was higher than 50% (I2=55%) in the allelic gene model, the heterogeneity was small in other genetic models.
The possibility of bias in our study was as follows. First, only English or Chinese-language studies were included in this meta-analysis, which might lead to potential publication bias, although publication bias was not significant in this study. Second, the exclusion of unpublished data was generally associated with an overestimation of the true effect. Third, a single population may not fully reflect the overall picture of other ethnicities. In addition, our study had other limitations. The sample number of the research objects included in the presented study was not large. Cancer risks can be regulated by factors, such as the interaction between genes and the environment and even the interaction between different polymorphic loci of the same gene. Further studies with larger samples that include different tumor types and ethnicities, are warranted, especially those studies involving interactions between genes or between genes and the environment should be given additional attention. Together, our analysis contributed to fully understanding the association of the MKK4 −1304T>G polymorphism with the risk of CRC.
Acknowledgements
Funding: This study was supported in part by grants from the Chinese National Natural Science Foundation (Grant Nos. 81572967, 81372498, and 81800429), the Hubei Natural Science Foundation (Grant No. 2013CFA006), the Zhongnan Hospital of Wuhan University Science, Technology and Innovation Seed Fund (Grant Nos. znpy2016050, znpy2017001, and znpy2017049), the National Key Clinical Specialty Construction Program of China (No. [2013]544), Wuhan City Huanghe Talents Plan and the Fundamental Research Funds for the Central Universities (Grant No. 2042018kf0065).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87-108. [Crossref] [PubMed]
- Jemal A, Bray F, Center MM, et al. Global cancer statistics. CA Cancer J Clin 2011;61:69-90. [Crossref] [PubMed]
- Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin 2016;66:115-32. [Crossref] [PubMed]
- Sohrabi M, Gholami A, Azar MH, et al. Trace Element and Heavy Metal Levels in Colorectal Cancer: Comparison Between Cancerous and Non-cancerous Tissues. Biol Trace Elem Res 2018;183:1-8. [Crossref] [PubMed]
- Wei Y, Wang L, Lan P, et al. The association between −1304T>G polymorphism in the promoter of MKK4 gene and the risk of sporadic colorectal cancer in southern Chinese population. Int J Cancer 2009;125:1876-83. [Crossref] [PubMed]
- Hu M, Zheng J, Zhang L, et al. The association between −1304T>G polymorphism in the promoter of mitogen-activated protein kinase kinase 4 gene and the risk of cervical cancer in Chinese population. DNA Cell Biol 2012;31:1167-73. [Crossref] [PubMed]
- Markowitz SD, Bertagnolli MM. Molecular origins of cancer: Molecular basis of colorectal cancer. N Engl J Med 2009;361:2449-60. [Crossref] [PubMed]
- Hemminki K, Chen B. Familial risk for colorectal cancers are mainly due to heritable causes. Cancer Epidemiol Biomarkers Prev 2004;13:1253-6. [PubMed]
- Lichtenstein P, Holm NV, Verkasalo PK, et al. Environmental and heritable factors in the causation of cancer--analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med 2000;343:78-85. [Crossref] [PubMed]
- Benna C, Helfrich-Forster C, Rajendran S, et al. Genetic variation of clock genes and cancer risk: a field synopsis and meta-analysis. Oncotarget 2017;8:23978-95. [Crossref] [PubMed]
- Lai SM, Zhang KB, Uhler RJ, et al. Geographic variation in the incidence of colorectal cancer in the United States, 1998-2001. Cancer 2006;107:1172-80. [Crossref] [PubMed]
- Liang W.. Age, sex and the risk of grade-specific second primary colorectal cancer: evidence for the protective effect of female hormone. Eur J Cancer 2007;43:1856-61. [Crossref] [PubMed]
- Mizoue T, Tanaka K, Tsuji I, et al. Alcohol drinking and colorectal cancer risk: an evaluation based on a systematic review of epidemiologic evidence among the Japanese population. Jpn J Clin Oncol 2006;36:582-97. [Crossref] [PubMed]
- Cho E, Smith-Warner SA, Ritz J, et al. Alcohol intake and colorectal cancer: a pooled analysis of 8 cohort studies. Ann Intern Med 2004;140:603-13. [Crossref] [PubMed]
- Misra UK, Pizzo SV. Ligation of cell surface GRP78 with antibody directed against the COOH-terminal domain of GRP78 suppresses Ras/MAPK and PI 3-kinase/AKT signaling while promoting caspase activation in human prostate cancer cells. Cancer Biol Ther 2010;9:142-52. [Crossref] [PubMed]
- Kim JS, Kim EJ, Kim HS, et al. MKK4 activates non-canonical NFkappaB signaling by promoting NFkappaB2-p100 processing. Biochem Biophys Res Commun 2017;491:337-42. [Crossref] [PubMed]
- Iqbal B, Masood A, Lone MM, et al. Polymorphism of Metastasis Suppressor Genes MKK4 and NME1 in Kashmiri Patients with Breast Cancer. Breast J 2016;22:673-7. [Crossref] [PubMed]
- Cunningham SC, Gallmeier E, Hucl T, et al. Targeted deletion of MKK4 in cancer cells: a detrimental phenotype manifests as decreased experimental metastasis and suggests a counterweight to the evolution of tumor-suppressor loss. Cancer Res 2006;66:5560-4. [Crossref] [PubMed]
- Wang L, Pan Y, Dai JL. Evidence of MKK4 pro-oncogenic activity in breast and pancreatic tumors. Oncogene 2004;23:5978-85. [Crossref] [PubMed]
- Yamada SD, Hickson JA, Hrobowski Y, et al. Mitogen-activated protein kinase kinase 4 (MKK4) acts as a metastasis suppressor gene in human ovarian carcinoma. Cancer Res 2002;62:6717-23. [PubMed]
- Yoshida BA, Dubauskas Z, Chekmareva MA, et al. Mitogen-activated protein kinase kinase 4/stress-activated protein/Erk kinase 1 (MKK4/SEK1), a prostate cancer metastasis suppressor gene encoded by human chromosome 17. Cancer Res 1999;59:5483-7. [PubMed]
- Nakayama K, Nakayama N, Davidson B, et al. Homozygous deletion of MKK4 in ovarian serous carcinoma. Cancer Biol Ther 2006;5:630-4. [Crossref] [PubMed]
- Davies H, Hunter C, Smith R, et al. Somatic mutations of the protein kinase gene family in human lung cancer. Cancer Res 2005;65:7591-5. [Crossref] [PubMed]
- Parsons DW, Wang TL, Samuels Y, et al. Colorectal cancer: mutations in a signalling pathway. Nature 2005;436:792. [Crossref] [PubMed]
- Lee JW, Soung YH, Kim SY, et al. Kinase domain mutation of MAP2K4 is rare in gastric, colorectal and lung carcinomas. Pathology 2006;38:263-4. [Crossref] [PubMed]
- Wang RF, Cheng Zhong, Lou LX. Association between MKK4 gene−1304T>G single nucleotide polymorphism and colorectal cancer. Zhongguo Lin Chuang Yao Li Xue Za Zhi 2015;31:615-7.
- Luo HL, Jun Huang, Bin Lai, et al. Effect of mkk4 gene promoter region polymorphism on susceptibility to colon cancer. Guangdong Yi Xue 2011;32:3208-11.
- Qin S. Association between the polymorphism of MKK4 gene promoter region and colorectal cancer. Hebei Medical University, 2014.
- Burotto M, Chiou VL, Lee JM, et al. The MAPK pathway across different malignancies: a new perspective. Cancer 2014;120:3446-56. [Crossref] [PubMed]
- Benhamman R, Bai F, Drory SB, et al. The Arabidopsis Mitogen-Activated Protein Kinase Kinase Kinase 20 (MKKK20) Acts Upstream of MKK3 and MPK18 in Two Separate Signaling Pathways Involved in Root Microtubule Functions. Front Plant Sci 2017;8:1352. [Crossref] [PubMed]
- Sun Y, Liu WZ, Liu T, et al. Signaling pathway of MAPK/ERK in cell proliferation, differentiation, migration, senescence and apoptosis. J Recept Signal Transduct Res 2015;35:600-4. [Crossref] [PubMed]
- Wagner EF, Nebreda AR. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat Rev Cancer 2009;9:537-49. [Crossref] [PubMed]
- Dong YQ, Lu CW, Zhang L, et al. Toll-like receptor 4 signaling promotes invasion of hepatocellular carcinoma cells through MKK4/JNK pathway. Mol Immunol 2015;68:671-83. [Crossref] [PubMed]
- Liu B, Chen D, Yang L, et al. A functional variant (−1304T>G) in the MKK4 promoter contributes to a decreased risk of lung cancer by increasing the promoter activity. Carcinogenesis 2010;31:1405-11. [Crossref] [PubMed]
- Ishikawa M, Nakayama K, Rahman MT, et al. Functional and clinicopathological analysis of loss of MKK4 expression in endometrial cancer. Oncology 2010;79:238-46. [Crossref] [PubMed]
- Su GH, Hilgers W, Shekher MC, et al. Alterations in pancreatic, biliary, and breast carcinomas support MKK4 as a genetically targeted tumor suppressor gene. Cancer Res 1998;58:2339-42. [PubMed]
- Cunningham SC, Kamangar F, Kim MP, et al. Claudin-4, mitogen-activated protein kinase kinase 4, and stratifin are markers of gastric adenocarcinoma precursor lesions. Cancer Epidemiol Biomarkers Prev 2006;15:281-7. [Crossref] [PubMed]
- Szmulewitz RZ, Clark R, Lotan T, et al. MKK4 suppresses metastatic colonization by multiple highly metastatic prostate cancer cell lines through a transient impairment in cell cycle progression. Int J Cancer 2012;130:509-20. [Crossref] [PubMed]
- Zheng J, Liu B, Zhang L, et al. The protective role of polymorphism MKK4-1304 T>G in nasopharyngeal carcinoma is modulated by Epstein-Barr virus' infection status. Int J Cancer 2012;130:1981-90. [Crossref] [PubMed]