Circulating microRNAs as emerging non-invasive biomarkers for gliomas
Introduction
Malignant gliomas are the most lethal primary brain tumors (1), conventionally classified by histopathologic evaluation into four tumor grades (I–IV), where glioblastoma (GBM), classified as grade IV glioma, is the most common and malignant. Even with the current standard treatment, consisting of maximal surgical resection and combined radio-chemotherapy (2), the prognosis for patients with GBM IV is very poor, with a median survival approximating 15 months, and only 3–5% of patients surviving longer than 36 months (3). The clinical management of patients with gliomas is often hindered by the lack of therapeutically convenient monitoring tools.
Computed tomography (CT) and magnetic resonance imaging (MRI) are the conventional techniques for disease staging and monitoring of tumor growth and response to therapy. However, both diagnosis and accurate grading of gliomas is often difficult or even impossible before surgery, because different pathological conditions such as metastasis from melanoma or primary tumors of the lung may have similar morphological patterns on MRI. In addition, pseudoprogression, a common finding related to the effect of radiotherapy treatment mimicking tumor recurrence, may additionally complicate the interpretation of imaging studies (4).
All these confounders clearly highlight the need for establishing reliable minimally invasive biomarkers to support and facilitate a more reliable and consistent diagnosis, staging and prognosis of patients affected by malignant gliomas.
Circulating biomarkers for gliomas
Circulating biomarkers are measurable biological molecules found in the blood or other body fluids, providing information about a particular condition, such as cancer. For brain tumors, circulating biomarkers are regarded as useful and easy accessible molecules that can be exploited for diagnostic purposes, especially in cases in which surgery is contraindicated or when biopsy results are inconclusive.
The most extensively studied circulating biomarkers are protein in nature, which can be actively or passively secreted by tumor cells and/or their microenvironment, and may be detected in blood, urine and cerebrospinal fluid (CSF). Many promising molecules have already been found in the blood and CSF from patients with gliomas (5), but none of these have yet entered routine practice. One of the few specific proteins in gliomas is the mutant EGFRvIII protein which, however, is not representative of all tumour cells. Correlative studies to date have shown that it could be a useful biomarker, but the clinical value of EGFRvIII is still pending validation. Glial fibrillary acidic protein (GFAP), a marker for circulating tumour cells has been also investigated as biomarker related to tumor burden (5).
Since proteins lack sufficient sensitivity and specificity needed for a successful biomarker, significant efforts have been made towards finding informative as well as non-invasive biomarkers for patients with gliomas.
The focus of this review will be circulating microRNA (miRNA) biomarkers for gliomas and how they can be further researched to support the clinical management of these lethal tumors.
miRNAs
miRNAs are endogenous, non-coding RNA molecules, of 18–22 nucleotides length, capable of modulating gene expression at the post-transcriptional level (6). As a result, under normal physiological conditions, miRNAs regulate almost all known biological processes including cell proliferation and differentiation, as well as cell metabolism and development (6). Regulation of gene expression by miRNAs is accomplished through a combination of translational repression and mRNA destabilization. Moreover, an intricate interplay between translational repression and mRNA degradation is emerging (6).
Due to their pivotal role in many biological processes, deregulation of miRNA expression interplays with cancer initiation and progression, thus exerting an impact on tumor behavior (7). As a matter of fact, miRNAs either have tumor suppressor capability or display oncogenic characteristics, both of which influence cell growth. Therefore, changes in a single or small subset of miRNAs have a profound effect on expression pattern of several hundred mRNAs, leading cells towards transformation (7).
The expression profiling of miRNAs has already entered cancer clinics as diagnostic and prognostic biomarkers to assess tumor initiation, progression and response to treatment. Evidence is also emerging suggesting that inhibition of oncogenic miRNAs could be used to develop cancer therapeutics, making them highly reliable and innovative biotargets (8).
miRNA expression profiles in gliomas
In addition to histology, the 2016 update of the World Health Organization (WHO) classification of tumors of the central nervous system (CNS) proposes molecular parameters, including miRNA detection, for improving both diagnostic accuracy and patient management (9). Large-scale miRNA expression analyses reported both the up-regulation and down-regulation of several miRNAs in tumour tissues from patients with GBM IV compared to normal brain tissue, which could hence permit a more reliable and consistent diagnosis (10). However, due to inconsistencies in the reported subset of miRNAs that are either up or down-regulated in GBM IV, a specific miRNA signature is not yet well established. Notably, miR-21 is the only miRNA found to be up-regulated in tumor tissues from GBM IV across all the studies, whereas among the down-regulated miRNAs, miR-132 is the most consistent, although it was identified only in 60% of the reported analyses (10). The quality of tumor tissues, sample sizes, appropriateness of “control” brain tissue and analytical techniques could explain discrepant results obtained across the published studies. Therefore, the potential of miRNAs as biomarkers for gliomas requires a systematic analysis of existing data. Recently, a meta-analysis of five studies published between 2012 and 2015, found 6 miRNAs differentially expressed, with a more accurate predictive value (i.e., higher sensitivity, specificity and statistical significance) in distinguishing glioma from control tissue (11). The miR-15a, miR-16, miR-21, miR-23a, miR-9 and miR-124 identified in this study may be useful diagnostic and prognostic biomarkers in glioma.
Status of circulating miRNA biomarkers
The discovery of miRNAs in tissues, as well as in body fluids together with their altered profile in various pathological conditions, offers a new perspective on the use of extracellular miRNAs as informative disease biomarkers. Circulating miRNAs have been the focus of numerous cancer biomarker identification efforts over the past few years, although most miRNA biomarkers reported in the literature have failed to enter clinical practice because of inconsistent and irreproducible findings. Before circulating miRNAs can reliably be used as biomarkers of disease, analytical factors that may affect their measurement, so including sample type, measurement platform or normalization strategy need to be addressed.
Major pre-analytical variables affecting circulating miRNAs have been characterized (12). Mechanical hemolysis, caused by improper blood collection or preparation, could alter the specificity of circulating miRNA expression because of contamination by intracellular miRNAs. Therefore, measurement miR-451, highly expressed in red blood cells, is recommended, in order to exclude samples with even low hemolysis that cannot be revealed visually (12).
Widespread technologies used to measure miRNA expression in biological samples include microarray, next generation sequencing (NGS), and quantitative real-time PCR (qPCR). Microarray-based miRNA measurement platforms provide overall miRNA expression profiles with reasonable cost and throughput. Nevertheless, although qPCR is more expensive per sample and has a lower throughput, the amplification technology offers a much higher sensitivity than microarrays (13). NGS can identify new miRNAs, is cheaper than microarray or qPCR and requires small amount of samples even though is tedious and has very long handling time. Therefore, qPCR remains the top choice for validation and clinical analysis of large numbers of samples (13).
Regardless of the platform used for miRNA measurement, data normalization to correct for variability during sample preparation and to quantify miRNA expression is another major issue. Addition of spike-in standards prior to cDNA synthesis and PCR can estimate efficiency and normalise results for comparison (14). The measurement of miRNA expression by digital PCR could circumvent normalization, being an absolute method of nucleic-acid quantification. In this respect, droplet digital PCR (ddPCR), a highly precise method and not reliant on reference standard curves, has recently been shown to reduce analytic variability and to improve day-to-day reproducibility compared to RT-qPCR (14).
miRNAs packaged in extracellular vesicles (EVs)
Plasma and serum carry EVs which are small lipid membrane-bound satchels expelled by cells to mediate cell-to-cell communication and containing soluble materials such as nucleic acids, lipids and proteins, protected from degradation (15). Emerging evidence suggests that exosomes, small EVs (40–150 nm) with a multivesicular endosomal origin secreted by both normal and neoplastic cells, play a crucial role in facilitating tumorigenesis (16). To prevent degradation in the circulation, miRNAs are released by cells in both exosomes and miRNA/protein complexes. Exosomes released from GBM IV have been for the first time demonstrated in serum of GBM IV patients by Skog and colleagues in 2008 (17), thus making it likely that changes relative to tumor progression can be detected using serum samples drawn over time.
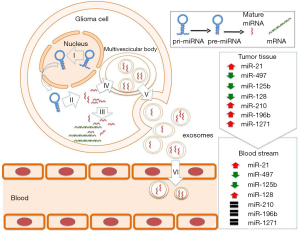
Both non-cancerous and cancerous cells release exosomes as a form of cellular communication and perform a variety of functions depending on their contents and cellular context. Although it is unclear as to how miRNAs are packaged into exosomes, this is a specific and selective process driven by the binding of sumoylated heterogeneous nuclear ribonucleoprotein A2/B1 (hnRNPA2B1) to specific motifs present in the sequence of mature miRNAs (18). This means that only certain miRNAs are incorporated and released into the circulation and, as a consequence, deregulated miRNAs within tumor cells may not be present within exosomes. As shown in Figure 1, miR-21 (which is known to be overexpressed in GBM IV tissue) was found to be elevated also in serum exosomes from patients with GBM IV (17). Other examples of circulating miRNAs as mirrors of tissue miRNA level in gliomas are miR-497 and miR-125b that have been described to be down-regulated both in serum and tissue tumors of patients with GBM IV (19). Unlike these biomarkers, no correlation between blood and tissue expression levels was reported for miR-128, which was found significantly increased in blood of GBM IV patients and down-regulated in GBM IV tissue (20). Moreover, miR-210, miR-196b and miR-1271 were found to be increased in brain tumors, but similar to healthy subjects in blood (21). Therefore, circulating miRNA signatures in exosomes need to be investigated independently of those for tumor tissues (22).
EVs can be isolated from body fluids via various methods based on their physical and chemical properties, resulting in different yield, purity and integrity of the final preparation. Current approaches, such as centrifugation, ultracentrifugation, filtration or affinity purification are very hands on, time consuming and not feasible in routine clinical practice (23). There are also precipitation-based methods which are fast, simple and economical, although they do not guarantee sufficient purity. Widely used chemical precipitation solutions, such as ExoquickTM from System Biosciences (Mountain View, CA, USA) and Total Exosome Isolation reagents from Life Technologies (Carlsbad, CA, USA) enable simple and efficient concentration of exosome-like EVs from small amounts of serum or plasma, although yield heterogeneous vesicular population (23). Sonication of blood or plasma is another possible approach, but has not been reliably tested so far.
Considering that circulating exosomes can be obtained by minimally invasive procedures and easily isolated from peripheral blood of patients with gliomas, a “liquid biopsy” based on miRNAs packaged in exosomes may be a practical alternative to tumor biopsy, that may not reflect the heterogeneity of gliomas, and would permit real-time assessment for monitoring recurrence and patient response to treatments, through longitudinal sampling along the course of the disease.
Exosome miRNAs as biomarkers for gliomas
Intriguing evidence showing an increase of EV number associated with poor prognosis, suggest that EV kinetics may be valuable in management of patients with GBM IV (24). Nevertheless, few studies investigated exosome miRNA expression in serum of patients with glioma so far.
MiR-21 is an abundantly expressed miRNA in mammalian cells, whose upregulation is associated with numerous types of cancer. MiR-21 functions as an anti-apoptotic and pro-survival factor, so that high levels of this miRNA may be a fingerprint of cancer cells and represent a common feature of pathological cell growth or cell stress (25). As mentioned in previous section, miR-21 is one of the most efficient clinical biomarker for diagnosing GBM IV (11). As far as exosome miRNA expression, miR-21 was found to be overexpressed in serum EVs from patients with GBM IV, and reduced after extensive resection of the tumor (17). Notably, levels of exosome miR-21 in CSF allow to discriminate GBM IV from non-oncologic patients (26) and circulating serum levels probably reflect the antiangiogenic effect of therapy with bevacizumab (21). Therefore, exosome-associated miR-21 expression may serve as a platform for glioma biomarker development.
Besides miR-21, a small miRNA signature has been found in serum exosomes of patients with GBM IV (27). The expression levels of small noncoding RNU6-1, miR-320 and miR-574-3p were observed to be significantly associated with a diagnosis of GBM IV. In addition, RNU6-1 was consistently an independent predictor for diagnosing GBM IV (27).
While these investigations of miRNA expression in EVs as a biomarker platform for gliomas show promise, validation through larger sample sets and prospective clinical trials are the necessary next steps.
Proposed circulating miRNAs as biomarkers for gliomas
As described above, the pool of circulating miRNAs contains two types of miRNAs: vesicle-packaged miRNAs and miRNAs complexed with the Argonaute-2 proteins, representing cell by-products that accumulate in extracellular fluids (28). Most of the studies on circulating miRNAs as biomarkers for gliomas focused on the entire circulating pool of miRNAs instead of vesicle-packaged miRNAs, so far.
Using comprehensive high-throughput approaches, such as microarrays, different miRNAs were reported as potential biomarkers of gliomas. However, most of these findings were not validated in other cohorts (22). Several patterns of altered expression profile of circulating miRNAs that may serve as biomarkers for tumor diagnosis and predict outcomes or the therapeutic response of patients with gliomas have been proposed. Some of the results obtained in different studies are summarized in Table 1.

Full table
The plasma levels of miR-21 (up-regulated), miR-128 and miR-342-3p (down-regulated) were found significantly altered in GBM IV patients compared to healthy controls. Notably, miR-128 and miR-342-3p positively correlated with histopathological grades of glioma (31). Down-regulation of miR-342-3p was also observed in another investigation whereas, on the contrary, miR-128 was found to be up-regulated in blood samples from GBM IV patients (20).
In a different study, combined expression analyses of miR-21 and miR-15b distinguished between patients with and without glioma (29).
A new additional tool to better characterize gliomas has been recently proposed by Zhang and colleagues (35), reporting plasma miR-221/222 family levels significantly up-regulated in glioma patients. Interestingly high plasma levels of miR-221/222 were associated with a lower survival rate.
Circulating miRNA expression has been suggested to predict tumor grading, as shown by up-regulation of serum miR-20a, miR-106a and miR-181b associated with advanced clinical stages of astrocytoma (30) or by miR-210, considerably increased in GBM IV (34). Notably, serum levels of miR-210 did not differ from those in healthy controls in the study by Siegal and colleagues (21). The possibility to distinguish high grade from low grade gliomas was proposed also by the expression of serum miR-497 and miR-125b (19) and miR-29 (33), which were all down-regulated in GBM IV (19).
miRNAs could be used as a disease follow-up marker as demonstrated by decreased levels of serum miR-21 after surgical tumor resection (17,33) or by persistently high expression associated with tumor progression (33).
Examples reported in Table 1, clearly prove that discovering a both sensitive and specific miRNA signature for glioma is still challenging. As stated above, the lack of clinical miRNA biomarkers compared to the number identified in research is due to limitations in standardizing preanalytical features such as choice of sample type and processing, as well as analytical methods, all of which can impair the reproducibility of individual findings. Furthermore, extensive heterogeneity and variety of subtypes of gliomas, may account for variability across different studies.
Nevertheless, a meta-analysis performed by Qu and colleagues to assess the diagnostic value of miRNAs revealed miR-21 as the most powerful clinical biomarker in diagnosis of GBM IV (36).
Conclusions
Circulating biomarkers represent an exciting area of research that holds promise for better management of patients with gliomas. Due to their scarce specificity and limited value to inform on molecular profiles and tumor heterogeneity, proteins lack wide clinical utility. miRNAs hold the potential to provide sensitive and specific biomarker panels. Their tissue and disease specificity make them ideal candidates for diagnostic and prognostic indicators. Several promising circulating miRNAs identified in small studies require further evaluation and validation in larger-scale prospective studies before their implementation in routine clinical practice. Among these miRNAs, miR-21 is of particular relevance, being one of the most upregulated miRNAs in GBM IV.
Current limitations of effective clinical applications of miRNA biomarkers will be likely overcome by strict methodological standardization. The difficulty in identifying a specific miRNA biomarker for glioma lies partly in the complex heterogeneous nature of the cancer itself. Multiple mutations occurring during transformation and genomic changes between grades and sub-types of gliomas contribute to this heterogeneity. Using a group of biomarkers to detect a range of these characteristics will be ultimately more beneficial than using a single biomarker.
Exosomes secreted by tumour cells are emerging as a key component in the biogenesis of glioma, in addition to support tumour progression and represent a potential source of miRNAs biomarkers for diagnosis and tracking cancer progression in patients with gliomas. Further efforts of research focused on exosome-related miRNAs are needed to advance the development of clinically useful biomarkers of gliomas.
Acknowledgements
We are grateful to Giampietro Pinna, Mario Meglio, Francesco Sala, Antonio Nicolato (Neurosurgery Institute), Giuseppe Moretto (Department of Neurological Sciences), Giuseppe K. Ricciardi, Elisa F. Ciceri, Alberto Beltramello (Neuroradiology Unit) of the University Hospital of Verona, Italy, and Giuseppe Lombardi and Vittorina Zagonel (Istituto Oncologico Veneto, Padova, Italy) for helpful discussions. This study is supported by Brain Research Foundation, Verona, Italy.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Schwartzbaum JA, Fisher JL, Aldape KD, et al. Epidemiology and molecular pathology of glioma. Nat Clin Pract Neurol 2006;2:494-503. [Crossref] [PubMed]
- Mutter N, Stupp R. Temozolomide: a milestone in neuro-oncology and beyond? Expert Rev Anticancer Ther 2006;6:1187-204. [Crossref] [PubMed]
- Stupp R, Hegi ME, Mason WP, et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomized phase III study: 5-Year analysis of the EORTC-NCIC trial. Lancet Oncol 2009;10:459-66. [Crossref] [PubMed]
- Brandsma D, Stalpers L, Taal W, et al. Clinical features, mechanisms, and management of pseudoprogression in malignant gliomas. Lancet Oncol 2008;9:453-61. [Crossref] [PubMed]
- Westphal M, Lamszus K. Circulating biomarkers for gliomas. Nat Rev Neurol 2015;11:556-66. [Crossref] [PubMed]
- Jonas S, Izaurralde E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet 2015;16:421-33. [Crossref] [PubMed]
- Reddy KB. MicroRNA (miRNA) in cancer. Cancer Cell Int 2015;15:38-44. [Crossref] [PubMed]
- Catela Ivkovic T, Voss G, Cornella H, et al. microRNAs as cancer therapeutics: A step closer to clinical application. Cancer Lett 2017;17:30237-9. [PubMed]
- Louis DN, Perry A, Reifenberger G, et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 2016;131:803-20. [Crossref] [PubMed]
- Areeb Z, Stylli SS, Koldej R, et al. MicroRNA as potential biomarkers in Glioblastoma. J Neurooncol 2015;125:237-48. [Crossref] [PubMed]
- Ye X, Wei W, Zhang Z, et al. Identification of microRNAs associated with glioma diagnosis and prognosis. Oncotarget 2017;8:26394-403. [PubMed]
- MacLellan SA, MacAulay C, Lam S, et al. Pre-profiling factors influencing serum microRNA levels. BMC Clin Pathol 2014;14:27-38. [Crossref] [PubMed]
- He Y, Lin J, Kong D, et al. Current State of Circulating MicroRNAs as Cancer Biomarkers. Clin Chem 2015;61:1138-55. [Crossref] [PubMed]
- Chevillet JR, Lee J, Briggs HA, et al. Issues and Prospects of microRNA-Based Biomarkers in Blood and Other Body Fluids. Molecules 2014;19:6080-105. [Crossref] [PubMed]
- Kosaka N, Yoshioka Y, Fujita Y, et al. Versatile roles of extracellular vesicles in cancer. J Clin Invest 2016;126:1163-72. [Crossref] [PubMed]
- Kalluri R. The biology and function of exosomes in cancer. J Clin Invest 2016;126:1208-15. [Crossref] [PubMed]
- Skog J, Würdinger T, van Rijn S, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 2008;10:1470-6. [Crossref] [PubMed]
- Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun 2013;4:2980-90. [Crossref] [PubMed]
- Regazzo G, Terrenato I, Spagnuolo M, et al. A restricted signature of serum miRNAs distinguishes glioblastoma from lower grade gliomas. J Exp Clin Cancer Res 2016;35:124-35. [Crossref] [PubMed]
- Roth P, Wischhusen J, Happold C, et al. A specific miRNA signature in the peripheral blood of glioblastoma patients. J Neurochem 2011;118:449-57. [Crossref] [PubMed]
- Siegal T, Charbit H, Paldor I, et al. Dynamics of circulating hypoxia-mediated miRNAs and tumor response in patients with high-grade glioma treated with bevacizumab. J Neurosurg 2016;125:1008-15. [Crossref] [PubMed]
- Tumilson CA, Lea RW, Alder JE, et al. Circulating MicroRNA biomarkers for glioma and predicting response to therapy. Mol Neurobiol 2014;50:545-58. [Crossref] [PubMed]
- Jia S, Zocco D, Samuels ML, et al. Emerging technologies in extracellular vesicle-based molecular diagnostics. Expert Rev Mol Diagn 2014;14:307-21. [Crossref] [PubMed]
- Evans SM, Putt M, Yang XY, et al. Initial evidence that blood-borne microvesicles are biomarkers for recurrence and survival in newly diagnosed glioblastoma patients. J Neurooncol 2016;127:391-400. [Crossref] [PubMed]
- Feng YH, Tsao CJ. Emerging role of microRNA-21 in cancer Biomed Rep 2016;5:395-402. (Review). [PubMed]
- Akers JC, Ramakrishnan V, Kim R, et al. miR-21 in the Extracellular Vesicles (EVs) of Cerebrospinal Fluid (CSF): A Platform for Glioblastoma Biomarker Development. PLoS One 2013;8:e78115. [Crossref] [PubMed]
- Manterola L, Guruceaga E, Perez-Larraya JG, et al. A small noncoding RNA signature found in exosomes of GBM patient serum as a diagnostic tool. Neuro-Oncol 2014;16:520-7. [Crossref] [PubMed]
- Turchinovich A, Weiz L, Langheinz A, et al. Characterization of extracellular circulating microRNA. Nucleic Acids Res 2011;39:7223-33. [Crossref] [PubMed]
- Ivo D’Urso P, Urso OF, Gianfreda CD. miR-15b and miR-21 as Circulating Biomarkers for Diagnosis of Glioma. Curr Genomics 2015;16:304-11. [Crossref] [PubMed]
- Zhi F, Shao N, Wang R, et al. Identification of 9 serum microRNAs as potential noninvasive biomarkers of human astrocytoma. Neuro-Oncol 2015;17:383-91. [Crossref] [PubMed]
- Wang Q, Li P, Li A, et al. Plasma specific miRNAs as predictive biomarkers for diagnosis and prognosis of glioma. J Exp Clin Cancer Res 2012;31:97-107. [Crossref] [PubMed]
- Ilhan-Mutlu A, Wagner L, Wohrer A, et al. Plasma MicroRNA-21 Concentration May Be a Useful Biomarker in Glioblastoma Patients. Cancer Invest 2012;30:615-21. [Crossref] [PubMed]
- Wu J, Li L, Jiang C. Identification and Evaluation of Serum MicroRNA-29 Family for Glioma Screening. Mol Neurobiol 2015;52:1540-6. [Crossref] [PubMed]
- Lai NS, Wu D, Fang X, et al. Serum microRNA-210 as a potential noninvasive biomarker for the diagnosis and prognosis of glioma. Br J Cancer 2015;112:1241-6. [Crossref] [PubMed]
- Zhang R, Pang B, Xin T, et al. Plasma miR-221/222 Family as Novel Descriptive and Prognostic Biomarkers for Glioma. Mol Neurobiol 2016;53:1452-60. [Crossref] [PubMed]
- Qu S, Guan J, Liu Y. Identification of microRNAs as novel biomarkers for glioma detection: a meta-analysis based on11 articles. J Neurol Sci 2015;348:181-7. [Crossref] [PubMed]


